6 附录
6.1.1常用抗生素配方
6.1.2常用母液配方
6.1.3缓冲液(pH 6-10)的pKa及缓冲范围
6.1.4磷酸缓冲液配方
6.2.1常用网站
6.2.2通用测序引物
6.2.3密码子及其偏好性表格
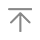
6.2.4空间群表格
6.2.5线站新衍射仪BL17U1
6.2.6实验室载体
6.2.7实验室细胞株及菌株
6.2.8结晶试剂盒
抗生素 | 母液浓度(mg/mL) | 工作浓度(μg/mL)for 严紧型质粒 | 工作浓度(μg/mL)for 松驰型质粒 | 处理方式 | |
Ampicillin,氨苄青霉素(Amp) | 100(溶于水) | 20 | 60 | 过滤除菌 | |
Kanamycin,卡那霉素(Kan) | 50 (溶于水) | 10 | 50 | 过滤除菌/高压 | |
Chloramphenicol,氯霉素(Chl/Cm/CAP) | 34 (溶于乙醇)
| 25 | 170 | 无需灭菌 | |
Carbenicillin,羧苄青霉素(Cab) | 50 (溶于水) | 20 | 60 | 过滤除菌 | |
Streptomycin,链霉素(Sm) | 10 (溶于水) | 10 | 50 | 过滤除菌 | |
Tetracyyline,四环素(Tet) | 5 (溶于甲醇) | 10 | 50 | 无需灭菌 | |
1. 保存条件:建议 -20 °C保存。
2. mycin,霉素,在不严格的缩写中作为“XX霉素”的后缀缩写为m。
3. 氯霉素(Chlor Am Phenicol),氯氨苯醇,标准缩写CAP,有时也写做Cm。
4. 过滤灭菌:用0.22 μm一次性滤膜过滤除菌,于超净台内。
5. 母液配制方法举例,如100 mg/mL氨苄青霉素:溶解1 g氨苄青霉素钠盐于足量的水中,最后定容至10 mL。分装成小份于-20 °C贮存。常以25 μg/mL~50 μg/mL的终浓度添加于生长培养基
1. 50× TAE(1 L):
Tris 242 g
Glacial Acetic Acid (冰醋酸) 57.1 mL
0.5 M EDTA 100 mL
2. 10× TBE(1 L):
Tris 108.8 g
Boric Acid 55.0 g
0.5 M EDTA 40 mL
3. 6×DNA Loading Buffer(100 mL):
1 M Tris-HCl pH 7.4 10 mL
100% Glycerol 60 mL
0.5 M EDTA 12 mL
Bromophenol Blue 0.01 g
4. 0.5 M EDTA pH 8.0(500 mL):
EDTA Disodium Salt Dihydrate 93.05 g
Dissolve with about 700 mL ddH2O, stir, then adjust pH to 8.0 (1 M NaOH)
Adjust the volume to 1 L. Filtered with 0.22 μm membrane or autoclave.
**EDTA does not dissolve until nearly pH 8.0
5. X-Gal (40 mg/mL), 1000×(5 mL)
X-Gal 0.200 g
Dimethyl Foramide 5 mL
(use glass pipette!)
Make ~500 μL aliquots. Store in brown glass vials at -20 °C.
6. SOC medium(250 mL)
Tryptone Peptone 5 g
Yeast extract 1.25 g
NaCl 0.15 g
KCl(10 mM) 0.05 g
MgCl2(10 mM) 0.51 g
50% glucose 4 mL
add sterile and MgCl2 and glucose after autoclaving
7. Solutions for Plasmid Preparations
1) Qiagen Buffer P1:50 mM Tris-HCl pH 8.0,10 mM EDTA,100 μg/mL RNaseA
&:for resuspension of cells in DNA plasmid preparation. 和生工P1通用。
1 M Tris pH 8.0 5 mL
0.5 M EDTA pH 8.0 2 mL
add ddH2O to 100 mL, Autoclave. Rnase A (add after autoclaving) 10.0 mg. Store at 4 °C
2) Qiagen Buffer P2:200 mM NaOH,1% SDS
&:for lysis of cells in DNA plasmid preparation. 和生工P2通用。
NaOH 0.8 g
SDS 1.0 g
add ddH2O to 100 mL
3) Qiagen Buffer P3:3.0 M potassium acetate pH 5.5,
&:not for spin columns, but for Qiatips, midi, maxi, giga kits
Potassium Acetate 25.03 g
Acetic Acid, Galacial ~11.5 mL
Adjust pH to 5.5, add ddH2O to 100 mL
4) Qiagen Buffer N3:4.2 M Gu-HCl,0.9 M potassium acetate pH 4.8
&:as nutrealization buffer to precipitate proteins and genomic materials,和生工小抽试剂盒P3通用。
5) Qiagen Buffer PB(5 M Gu-HCl, 30% isopropanol)=PBI(I for pH indicator)
6) Qiagen Buffer QG(5.5 M guanidine thiocyanate (GuSCN),20 mM Tris HCl pH 6.6)
7) Qiagen Buffer PE(10 mM Tris-HCl pH 7.5,80% ethanol )
&:for spin column salt wash in miniprep (blue)or pink columns. 用于质粒小抽和核酸提取离心柱清洗除盐。
8) Qiagen Buffer QX1:7 M NaPO4,10 mM NaAc pH 5.3
&:for solution and binding of agarose gels,用在用琼脂糖珠子DNA纯化试剂盒。
9) Qiagen Buffer QXB:5 M GuHCl
&:for binding of large >3000 bp fragments to columns
10) Qiagen Buffer QBT:750 mM NaCl,50 mM MOPS pH 7.0,15% isopropanol,0.15% triton X-100
&:Equilibration buffer
NaCl 4.38 g
Isopropanol 15 mL
MOPS 1.05 g
Triton X-100 150 mL
Adjust pH to 7.0(use 1N), add ddH2O to 100 mL
11) Qiagen Buffer QC:1.0 M NaCl,50 mM MOPS pH 7.0,15% isopropanol
&:Wash buffer
NaCl 5.84 g
MOPS 1.05 g
Isopropanol 15 mL
Adjust pH to 7.0, add ddH2O to 100 mL ,filter sterilize.
12) Qiagen Buffer QF:1.25 M NaCl, 50 mM Tris-HCl pH 8.5,15% isopropanol
&:Elution buffer
NaCl 7.31 g
Tris 8.5 1 M 5 mL
Isopropanol 15 mL
Adjust pH to 8.5, add ddH2O to 100 mL
8. Buffers for Protein Electrophoresis
1) 6×SDS Sample Buffer
1 M Tris-HCl pH 6.8 25 mL
10% SDS 20 mL
50% Glycerol 50 mL
1 M DTT 2 mL
Bromophenol Blue 0.01 g
Add ddH2O to 100 mL
2) Coommassie Blue Staining Solution(500 mL):
Coommassie Brilliant Blue R250 0.5 g
Ethanol 200 mL
Acetic acid: 50 mL
ddH2O to 500 mL(~250 mL)
(Dissolve Coommassie in Ethanol before adding water and acetic acid will require stirring 1~2 hours)
3) Coommassie Blue Destaining Solution (500 mL):
Ethanol 200 mL
Acetic Acid 50 mL
ddH2O 250 mL
4) 20×MES-SDS running buffer (1 L):MES:2-(N-吗啉)乙磺酸一水合物
&:For high-resolution separation of small size proteins.
MES 195.2 g
Tris 121.2 g
SDS 20 g
EDTA 6 g
9. Molecular Biology Buffers:
1) A1:1×NEBuffer 1(yellow):10 mM Bis Tris Propane-HCl,10 mM MgCl2,1 mM DTT pH 7.0 @ 25 °C。Supplied as a 10×concentrated stock.
2) A2:1×NEBuffer 2(blue):50 mM NaCl, 10 mM Tris-HCl,10 mM MgCl2,1 mM DTT pH 7.9 @ 25 °C。Supplied as a 10×concentrated stock.
3) A2:1×NEBuffer 3(red):100 mM NaCl, 50 mM Tris-HCl,10 mM MgCl2,1 mM DTT pH 7.9 @ 25 °C。Supplied as a 10×concentrated stock.
4) A2:1×NEBuffer 4(green,cutsmart buffer):50 mM potassium acetate,20 mM Tris acetate,10 mM magnesium acetate,1 mM DTT, pH 7.9 @ 25 °C。Supplied as a 10×concentrated stock.
5) 2×T4 DNA quick ligase 快连接酶buffer:60 mM Tris-HCl pH 7.8,20 mM MgCl2,2 mM DTT,2 mM ATP and 10% PEG6000
6) 10×T4 DNA连接酶buffer:500 mM Tris-HCl pH 7.6,100 mM MgCl2,100 mM DTT,10 mM ATP
7) 10×pfu buffer:200 mM Tris-HCl pH 8.6,100 mM KCl,160 mM (NH4)2SO4,10 mM MgCl2,1% Triton X-100,1 mg/mL BSA。
8) 5×HF buffer:32 mM HEPES-KOH buffer pH 7.8,100 mM KAcO, 4 mM Mg (AcO) 2,0.05% BSA(0.5 mg/mL)
9) 5×GC rich buffer:32 mM HEPES-KOH buffer pH 7.8,100 mM KAcO,7.5 mM Mg(AcO)2,0.01% BSA(0.1 mg/mL),1.0% DMSO
10) 2×Taq mix:
1 M Tris-HCl pH 8.6 4 mL
1 M KCl 20 mL
1 M MgCl2 600 μL
Sucrose 68 g
dNTP (2 mM each) 40 mL
1% Cresol Red(甲酚红) 2 mL
Taq polymerase (2U/μL) 20 mL
Add ddH2O to 200 mL
11) 10×Taq buffer:
200 mM Tris-HCl 8.4,200 mM KC,100 mM (NH4)2SO4,30 mM MgSO4,1 mg/mL BSA
6.1.3 缓冲液(pH 6-10)的pKa及缓冲范围
缓冲液 | pKa(25°C) | 缓冲范围 | 分子量 | 中文全称 |
MES | 6.1 | 5.5~6.7 | 195.2 | 2-吗啉乙磺酸 |
Bis-Tris | 6.5 | 5.8~7.2 | 209.2 | 二(2-羟乙基)亚胺基三(羟甲基)甲烷 |
ADA | 6.6 | 6.0~7.2 | 190.2 | N-(氨基甲酰基甲基)亚氨基二乙酸 |
ACES | 6.8 | 6.1~7.5 | 182.2 | N-氨基甲酰甲基乙磺酸 |
PIPES | 6.8 | 6.1~7.5 | 302.4 | 哌嗪-N,N'-二(2-乙磺酸) |
MOPSO | 6.9 | 6.2~7.6 | 225.3 | 3-(N-吗啉基)-2-羟基丙磺酸 |
Bis-Tris Propane | 6.8 | 6.3~9.5 | 282.3 | 双[三(羟甲基)氨基丙烷]/1,3-二[三(羟甲基)甲氨基]丙烷 |
BES | 7.1 | 6.4~7.8 | 213.2 | N-双(2-羟乙基)-2-氨基乙璜酸 |
MOPS | 7.2 | 6.5~7.9 | 209.3 | 3-(N-吗啡啉)乙磺酸 |
HEPES | 7.5 | 6.8~8.2 | 238.3 | N-2-羟乙基哌嗪-N'-2-乙磺酸 |
TES | 7.4 | 6.8~8.2 | 229.2 | N-3-(羟甲基)甲基-2-氨基乙磺酸 |
DIPSO | 7.6 | 7.0~8.2 | 243.3 | 3-[N-N-双(2-羟乙基)氨基]-2-羟基丙磺酸 |
TAPSO | 7.6 | 7.0~8.2 | 259.3 | N-3-(羟甲基)甲氨基-2-羟基丙烷磺酸 |
Tris | 8.1 | 7.0~9.1 | 121.1 | 三羟甲基氨基甲烷 |
HEPPSO | 7.8 | 7.1~8.5 | 268.3 | 4-(2-羟乙基)哌嗪-1-2-羟基丙磺酸 |
POPSO | 7.8 | 7.2~8.5 | 362.4 | 哌嗪-1,4-二羟基丙磺酸 |
EPPS | 8.0 | 7.3~8.7 | 252.3 | 4-羟乙基哌嗪丙磺酸 |
TEA | 7.8 | 7.3~8.3 | 149.2 | 三乙醇胺 |
Tricine | 8.1 | 7.4~8.8 | 179.2 | N-三-(羟甲基)甲基氨基乙酸 |
Bicine | 8.3 | 7.6~9.0 | 163.2 | N,N-双(2-羟乙基)甘氨酸 |
TAPS | 8.4 | 7.7~9.1 | 243.3 | 三羟甲基甲胺基丙磺酸 |
AMPSO | 9.0 | 8.3~9.7 | 227.3 | 3-[N-(1,1-二甲基-2-羟乙基)]氨基-2-羟丙烷磺酸 |
CHES | 9.3 | 8.6~10.0 | 207.3 | 2-(环已胺)-1-乙磺酸 |
CAPSO | 9.6 | 8.9~10.3 | 237.3 | 3-(环已胺)-2-羟基-1-丙磺酸 |
AMP | 9.7 | 9.0~10.5 | 89.1 | 2-氨基-2-甲基-1-丙醇 |
CAPS | 10.4 | 9.7~11.1 | 221.3 | 3-(环已胺)-1-丙磺酸 |
材料(MATERIALS)
r 试剂(REAGENTS)
NaH2PO4·2H2O、Na2HPO4·12H2O
配制方法:
配制时,常先配制0.2 M的NaH2PO4和0.2 M的Na2HPO4,两者按一定比例混合即成0.2 M的磷酸盐缓冲液(PB),根据需要可配制不同浓度的PB和PBS。
1. 0.2 M的 NaH2PO4:称取NaH2PO4.2H2O 31.21 g(或 NaH2PO4·H2O 27.6 g)加重蒸水至1000 mL溶解。
2. 0.2 M的 Na2HPO4:称取Na2HPO4·12H2O 71.64 g(或 Na2HPO4·7H2O 53.6 g或 Na2HPO4·2H2O 35.61 g)加重蒸水至1000 mL溶解。
3. 0.2 M pH 7.4的PB的配制:取19 mL 0.2 M的NaH2PO4和81 mL 0.2 M Na2HPO4,充分混合即为0.2 M的PB(pH约为7.4~7.5)。若pH偏高或偏低,可通过改变二者的比例来加以调整,室温保存即可。
► 25 °C下0.2 M磷酸钠缓冲液的配制
pH值 | 0.2 M Na2HPO4(mL) | 0.2 M NaH2PO4(mL) |
5.8 | 8.0 | 92.0 |
5.9 | 10.0 | 90.0 |
6.0 | 12.3 | 87.7 |
6.1 | 15.0 | 85.0 |
6.2 | 18.5 | 81.5 |
6.3 | 22.5 | 77.5 |
6.4 | 26.5 | 73.5 |
6.5 | 31.5 | 68.5 |
6.6 | 37.5 | 62.5 |
6.7 | 43.5 | 56.5 |
6.8 | 49.0 | 51.0 |
6.9 | 55.0 | 45.0 |
7.0 | 61.0 | 39.0 |
7.1 | 67.0 | 33.0 |
7.2 | 72.0 | 28.0 |
7.3 | 77.0 | 23.0 |
7.4 | 81.0 | 19.0 |
7.5 | 84.0 | 16.0 |
7.6 | 87.0 | 13.0 |
7.7 | 89.5 | 10.5 |
7.8 | 91.5 | 8.5 |
7.9 | 93.5 | 6.50 |
8.0 | 94.7 | 5.3 |
► 25 °C下0.1 M磷酸钾缓冲液的配制
pH值 | 1 M K2HPO4(mL) | 1 M KH2PO4(mL) |
5.8 | 8.5 | 91.5 |
6.0 | 13.2 | 86.8 |
6.2 | 19.2 | 80.8 |
6.4 | 27.8 | 72.2 |
6.6 | 38.1 | 61.9 |
6.8 | 49.7 | 50.3 |
7.0 | 61.5 | 38.5 |
7.2 | 71.7 | 28.3 |
7.4 | 80.2 | 19.8 |
7.6 | 86.6 | 13.4 |
7.8 | 90.8 | 9.2 |
8.0 | 94.0 | 6.0 |
1. 分子克隆(Molecular Cloning)
1) 引物Tm值计算:http://biotools.nubic.northwestern.edu/OligoCalc.html
2) 密码子优化:https://sg.idtdna.com/CodonOpt
3) 质粒图谱分析:http://wishart.biology.ualberta.ca/PlasMapper/
4) 启动子信息:https://blog.addgene.org/plasmids-101-the-promoter-region
5) addgene(一个质粒数据库):http://www.addgene.org/vector-database/
6) NEB 内切酶buffer推荐:https://nebcloner.neb.com/#!/redigest
7) 核酸数据分析:http://biotools.nubic.northwestern.edu/OligoCalc.html
2. 蛋白表达与纯化(Protein Expression and Purification)
1) 蛋白表达菌株信息:http://wolfson.huji.ac.il/expression/bac-strains-prot-exp.html
2) 蛋白纯化:http://wolfson.huji.ac.il/purification/Purification_Protocols.html
3. 蛋白序列分析(Protein Sequence Analysis)
1) 信号肽预测:http://www.cbs.dtu.dk/services/SignalP/
2) 跨膜区预:TMpred http://www.ch.embnet.org/software/TMPRED_form.html
3) 结构域预测:http://smart.embl-heidelberg.de/
4) 蛋白性质预测:http://web.expasy.org/protparam/
http://biotools.nubic.northwestern.edu/proteincalc.html
5) 蛋白质结构与功能预测:http://raptorx.uchicago.edu/
6) 二级结构预测:http://bioinf.cs.ucl.ac.uk/psipred/
7) 评估结构预测的准确性和可靠性:https://www.cameo3d.org/
8) 蛋白质结构和功能预测:https://open.predictprotein.org/
9) 三维结构比对http://ekhidna.biocenter.helsinki.fi/dali_server/start
10) 同源建模https://zhanglab.ccmb.med.umich.edu/I-TASSER/
11) 同源建模:https://swissmodel.expasy.org/
12) 预测蛋白相互作用:https://string-db.org/
13) Pfam数据库是蛋白质家族的集合(根据结构域分类):http://pfam.xfam.org/
4. 结构分析(Structure Analysis)
1) RCSB PDB 数据库:http://www.rcsb.org/
2) 晶体界面分析 PISA serverhttp://www.ebi.ac.uk/pdbe/pisa/
3) Deposition serverhttps://deposit-1.wwpdb.org/
4) 配体结构文件下载:http://www.rcsb.org/pdb/ligand/chemAdvSearch.do
5) Bernhard Rupp的生物大分子晶体学网站:http://www.ruppweb.org/Xray/101index.html
5. 其他
1) ATCC:http://www.attc.org/
2) 化学分子性质:http://www.chemicalbook.com/ProductIndex_EN.aspx
3) RNA修饰数据库:http://mods.rna.albany.edu/mods/modifications/
4) Wikipedia:https://www.wikipedia.org/
5) 各类protocols:http://openwetware.org/wiki/Protocols
6) 文献查询:https://sci-hub.tw/
6.2.2 通用测序引物
简介(INTRODUCTION)
当进行菌落PCR时,如果目的片段没有成功转化至细菌中,针对插入片段的特异性引物可能会与细菌自身DNA进行非特异性结合,产生非特异性条带,从而对我们鉴定克隆是否成功构建产生干扰。因此我们常用载体上的引物进行鉴定。利用这种引物,我们既可以检测转入的质粒是否是空载体,也能估测连入载体的插入片段大小是否正确。
以下是我们实验室常用的几个通用引物,既可以做菌落PCR,也可以用于测序。
Primer name | Site | Sequence |
Z5 | T7 promoter | TACGACTCACTATAGGGGAATTGTGAGCG |
Z6 | T7 terminator | |
Z8 | TEV and Sal I | gagaatctttatttccaaggttctgTCGAC |
Z9 | v52 Rev | GGCCAGTGAATTGTAATACGACTCACTATA |
Z10 | v55 Rev | GTTCTGATTTAATCTGTATCAGGCTGAAAA |
Z11 | pCold (v113, v114) Rev | aaatggcagggatcttagattctgtg |
XY27 | pcDNA3.1 For | taatacgactcactatagggagacccaagc |
XY28 | pcDNA3.1 Rev | tagaaggcacagtcgaggctgatca |
y936 | v108 Rev | ggctgattatgatcctctagtacttc |
Z12 | v104 For | aatttgtaatacgactcactatagggcga |
Z13 | v104 Rev | tttagaggccccaaggggttatgctagtt |
Z14 | v105 For | agatctttaatacgactcactatagggcga |
Z15 | v105 Rev | ttgagatggtgcacgatgcacagttgaa |
Z16 | V126, 127 For | agaacccactgcttactggcttatcgaaat |
Z17 | V126, 127 Rev | ctgatcagcgggtttaaactcaatggtgat |
L22 | pEGFP-N1 For | tctatataagcagagctggtttagtgaaccgtcag |
L23 | pEGFP-N1 Rev | tgcacgccgtaggtcagggtggtcac |
L28 | pCDH For | GAGCTCGTTTAGTGAACCGTCAGATCGCCT |
L29 | pCDH Rev | GTTCAATTGCCGACCCCTCCCCCCAACTTC |
第一位碱基 | 第二位碱基 | |||
U | C | A | G | |
U | UUU (Phe/F)苯丙氨酸 | UCU (Ser/S)丝氨酸 | UAU (Tyr/Y)酪氨酸 | UGU (Cys/C)半胱氨酸 |
UUC (Phe/F)苯丙氨酸 | UCC (Ser/S)丝氨酸 | UAC (Tyr/Y)酪氨酸 | UGC (Cys/C)半胱氨酸 | |
UUA (Leu/L)亮氨酸 | UCA (Ser/S)丝氨酸 | UAA (终止) | UGA (终止) | |
UUG (Leu/L)亮氨酸 | UCG (Ser/S)丝氨酸 | UAG (终止) | UGG (Trp/W)色氨酸 | |
C | CUU (Leu/L)亮氨酸 | CCU (Pro/P)脯氨酸 | CAU (His/H)组氨酸 | CGU (Arg/R)精氨酸 |
CUC (Leu/L)亮氨酸 | CCC (Pro/P)脯氨酸 | CAC (His/H)组氨酸 | CGC (Arg/R)精氨酸 | |
CUA (Leu/L)亮氨酸 | CCA (Pro/P)脯氨酸 | CAA (Gln/Q)谷氨酰胺 | CGA (Arg/R)精氨酸 | |
CUG (Leu/L)亮氨酸 | CCG (Pro/P)脯氨酸 | CAG (Gln/Q)谷氨酰胺 | CGG (Arg/R)精氨酸 | |
A | AUU (Ile/I)异亮氨酸 | ACU (Thr/T)苏氨酸 | AAU (Asn/N)天冬酰胺 | AGU (Ser/S)丝氨酸 |
AUC (Ile/I)异亮氨酸 | ACC (Thr/T)苏氨酸 | AAC (Asn/N)天冬酰胺 | AGC (Ser/S)丝氨酸 | |
AUA (Ile/I)异亮氨酸 | ACA (Thr/T)苏氨酸 | AAA (Lys/K)赖氨酸 | AGA (Arg/R)精氨酸 | |
AUG (Met/M)甲硫氨酸 | ACG (Thr/T)苏氨酸 | AAG (Lys/K)赖氨酸 | AGG (Arg/R)精氨酸 | |
G | GUU (Val/V)缬氨酸 | GCU (Ala/A)丙氨酸 | GAU (Asp/D)天冬氨酸 | GGU (Gly/G)甘氨酸 |
GUC (Val/V)缬氨酸 | GCC (Ala/A)丙氨酸 | GAC (Asp/D)天冬氨酸 | GGC (Gly/G)甘氨酸 | |
GUA (Val/V)缬氨酸 | GCA (Ala/A)丙氨酸 | GAA (Glu/E)谷氨酸 | GGA (Gly/G)甘氨酸 | |
GUG (Val/V)缬氨酸 | GCG (Ala/A)丙氨酸 | GAG (Glu/E)谷氨酸 | GGG (Gly/G)甘氨酸 | |
密码子偏好性表格:
人类密码子偏好性表格
第一位碱基 | 第二位碱基 | 第三位碱基 | |||
U | C | A | G | ||
U | UUU(17.6%) | UCU(15.2%) | UAU(12.2%) | UGU(10.6%) | U |
UUC(20.3%) | UCC (17.7%) | UAC(15.3%) | UGC (12.6%) | C | |
UUA(7.7%) | UCA(12.2%) | UAA(1%) | UGA (1.6%) | A | |
UUG(12.9%) | UCG(4.4%) | UAG (0.8%) | UGG (13.2%) | G | |
C
| CUU (13.2%) | CCU(17.5%) | CAU(10.9%) | CGU(4.5%) | U |
CUC (19.6%) | CCC(19.8%) | CAC(15.1%) | CGC(10.4%) | C | |
CUA (7.2%) | CCA(16.9%) | CAA(12.3%) | CGA(6.2%) | A | |
CUG(39.6%) | CCG(6.9%) | CAG(34.2%) | CGG(11.4%) | G | |
A | AUU(16%) | ACU (13.1%) | AAU(17%) | AGU(12.1%) | U |
AUC (20.8%) | ACC(18.9%) | AAC(19.1%) | AGC(19.5%) | C | |
AUA(7.5%) | ACA(15.1%) | AAA(24.4%) | AGA(12.2%) | A | |
AUG(22%) | ACG(6.1%) | AAG(31.9%) | AGG(12%) | G | |
G | GUU(11%) | GCU(18.4%) | GAU(21.8%) | GGU(10.8%) | U |
GUC(14.5%) | GCC(27.7%) | GAC(25.1%) | GGC(22.2%) | C | |
GUA (7.1%) | GCA(15.8%) | GAA(29%) | GGA(16.5%) | A | |
GUG (28.1%) | GCG(7.4%) | GAG(39.6%) | GGG(16.5%) | G | |
大肠杆菌密码子偏好性表格
第一位碱基 | 第二位碱基 | 第三位碱基 | |||
U | C | A | G | ||
U | UUU(24.4%) | UCU(13.1%) | UAU(21.6%) | UGU(5.9%) | U |
UUC(13.9%) | UCC (9.7%) | UAC(11.7%) | UGC (5.5%) | C | |
UUA(17.4%) | UCA(13.1%) | UAA(2%) | UGA (1.1%) | A | |
UUG(12.9%) | UCG(8.2%) | UAG (0.3%) | UGG (13.4%) | G | |
C
| CUU (14.5%) | CCU(9.5%) | CAU(12.4%) | CGU(15.9%) | U |
CUC(9.5%) | CCC(6.2%) | CAC(7.3%) | CGC(14%) | C | |
CUA (5.6%) | CCA(9.1%) | CAA(14.4%) | CGA(4.8%) | A | |
CUG(37.4%) | CCG(14.5%) | CAG(26.7%) | CGG(7.9%) | G | |
A | AUU(29.6%) | ACU (13.1%) | AAU(29.3%) | AGU(13.2%) | U |
AUC (19.4%) | ACC(18.9%) | AAC(20.3%) | AGC(14.3%) | C | |
AUA(13.3%) | ACA(15.1%) | AAA(37.2%) | AGA(7.1%) | A | |
AUG(23.7%) | ACG(13.6%) | AAG(15.3%) | AGG(4%) | G | |
G | GUU(21.6%) | GCU(18.9%) | GAU(33.7%) | GGU(23.7%) | U |
GUC(13.1%) | GCC(21.6%) | GAC(17.9%) | GGC(20.6%) | C | |
GUA (13.1%) | GCA(23%) | GAA(35.1%) | GGA(13.6%) | A | |
GUG (19.9%) | GCG(21.1%) | GAG(19.4%) | GGG(12.3%) | G | |
6.2.4 空间群表格
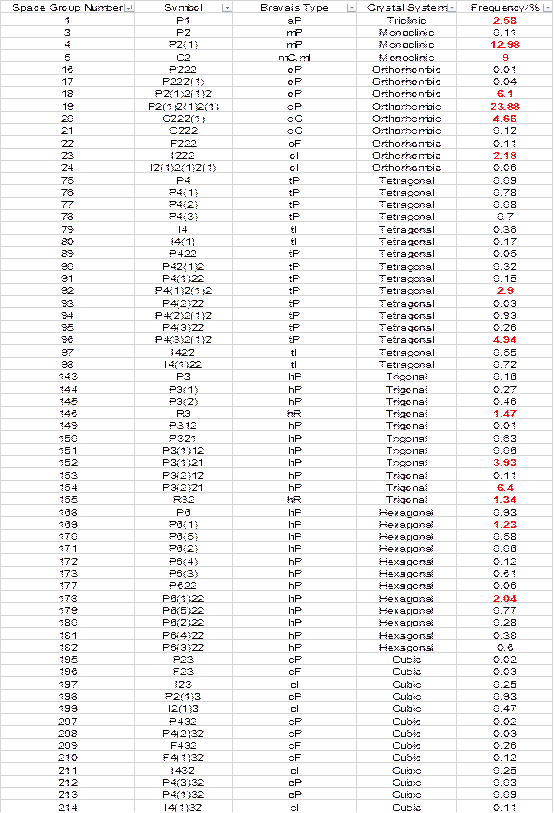
6.2.5 线站新衍射仪BL17U1
简介(INTRODUCTION)
上海光源BL17U1线站新衍射仪(自主研制,水平方向SOC~±0.342 μm,垂直方向SOC~±0.319 μm)和新探测器(瑞士Dectris公司的Eiger X 16 M)已经安装调试完成,11月10号开始正式对用户开放使用。但是在开放初期还需要各位老师提醒实验人员注意以下事项:
1. 由于衍射仪测角头电机比较小,如果手动上样的话很容易把位置碰歪,所以目前只能机械手上样,建议各位老师将晶体样品保存在puck里,如果没有puck,可以提前到线站借用我们线站的puck将晶体样品转移到puck里再通过机械手上样。
2. 如果在实验的过程中发生故障,一定要第一时间给值班人员打电话,不能自行去处理。
3. 建议在EIGER 16 M探测器上收数据的策略是:0.1~0.3度一张,曝光时间0.05秒一张。这样收一套360度的完整数据(3600张衍射图)大概只需要3分钟。
4. 一套3600张衍射图的数据大小是25 GB左右,建议带一个存储空间大的硬盘拷贝数据,可以边实验边拷贝数据。减少拷贝数据等待时间。
5. 数据处理建议使用XDS。
6. 目前线站的新系统还需要进一步优化调试,能量暂时固定在12.660 Kev。
Name | Backnone | Description | Cloning Frame | Fusion Tag | Sequencing Primer 1 | Sequencing Primer2 | Resistance |
|
|
V1 | pET30a | NdeI-SalI,NotI-XhoI-His6-stop | SalI-NotI | none | T7 Promoter | T7 Terminator | Kan |
|
|
V2 | pET30a | NdeI-His8-SalI,NotI-stop | SalI-NotI | none | T7 Promoter | T7 Terminator | Kan |
|
|
V3 | pET30a | NdeI-His8-GB1-TEV-SalI,NotI-stop | SalI-NotI | His8-GB1-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V4 | pET30a | NdeI-His8-GB1-TEV-SalI,NotI-XhoI-His6-stop | SalI-NotI | His8-GB1-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V5 | pET30a | NdeI-His8-TEV-SalI,NotI-stop | SalI-NotI | MGSS-His8-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V6 | pET30a | NdeI-His8-TEV-SalI,NotI-XhoI-His6-stop | SalI-NotI | MGSS-His8-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V7 | pET30a | NdeI-His8-SUMO-TEV-SalI,NotI-XhoI-stop | SalI-NotI | MGSS-His8-SUMO-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V8 | pET30a | NdeI-His8-VHb-TEV-SalI,NotI-stop | SalI-NotI | His8-VHb-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V9 | pET30a | NdeI-His8-VHb-TEV-SalI,NotI-XhoI-His6-stop | SalI-NotI | His8-VHb-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V10 | pET30a | NdeI-His8-GST-TEV-SalI,NotI-stop | SalI-NotI | His8-GST-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V11 | pET30a | NdeI-His8-GST-TEV-SalI,NotI-XhoI-His6-stop | SalI-NotI | His8-GST-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V12 | pET30a | NdeI-His7-TRX-TEV-SalI,NotI-stop | SalI-NotI | His7-TRX-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V13 | pET30a | NdeI-His7-TRX-TEV-SalI,NotI-XhoI-His6-stop | SalI-NotI | His7-TRX-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V14 | pET30a | NdeI-His8-MBP-SalI,NotI-stop | SalI-NotI | His8-MBP | T7 Promoter | T7 Terminator | Kan |
|
|
V15 | pET30a | NdeI-His8-tag2-TEV-SalI,NotI-stop | SalI-NotI | His8-tag2-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V16 | pET30a | NdeI-His8-tag2-TEV-SalI,NotI-XhoI-His6-stop | SalI-NotI | His8-tag2-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V17 | pET30a | NdeI-DsbA-TEV-SalI,NotI-XhoI-His6-stop | SalI-NotI | DsbA-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V18 | pET30a | NdeI-MBP-SalI,NotI-XhoI-His6-stop | SalI-NotI | MBP | T7 Promoter | T7 Terminator | Kan |
|
|
V19 | pET30a | NdeI-His8-NusA-TEV-SalI,NotI-stop | SalI-NotI | His8-NusA-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V20 | pET30a | NdeI-His8-GB1-TEV-SalI,NotI-Acidic Peptide-LacZ-XhoI-His6-stop | SalI-NotI | His8-GB1-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V21 | pET30a | NdeI-His8-MBP-helical linker-SalI,NotI-stop | SalI-NotI | His8-MBP-helical linker | T7 Promoter | T7 Terminator | Kan |
|
|
V22 | pMAL | NdeI-MBP-SalI,NotI-XhoI-LacZ-stop | SalI-NotI | MBP | M13R-pUC | M13F-pUC | Amp |
|
|
V23 | pMAL | NdeI-MBP-SalI,NotI-XhoI-LacZ-stop | SalI-NotI | MBP | M13R-pUC | M13F-pUC | Amp |
|
|
V24 | pMAL | NdeI-MBP-SalI,NotI-XhoI-LacZ-stop | SalI-NotI | MBP | M13R-pUC | M13F-pUC | Amp |
|
|
V25 | pMAL | NdeI-MBP-SalI,NotI-XhoI-LacZ-stop | SalI-NotI | MBP | M13R-pUC | M13F-pUC | Amp |
|
|
V26 | pMAL | NdeI-MBP-SalI,NotI-XhoI-LacZ-stop | SalI-NotI | MBP | M13R-pUC | M13F-pUC | Amp |
|
|
V27 | pMAL | NdeI-MBP-SalI,NotI-XhoI-LacZ-stop | SalI-NotI | MBP | M13R-pUC | M13F-pUC | Amp |
|
|
V28 | pET30a | NdeI-MBP-SalI,NotI-XhoI-His6-stop | SalI-NotI | MBP | T7 Promoter | T7 Terminator | Kan |
|
|
V29 | pET30a | NdeI-MBP-TEV-SalI,NotI-stop | SalI-NotI | MBP-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V29H | pET30a | NdeI-His8-MBP-TEV-SalI,NotI-stop | SalI-NotI | His8-MBP-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V30 | pET30a | NdeI-MBP-TEV-SalI,NotI-XhoI-His6-stop | SalI-NotI | MBP-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V31 | pLEXSY-Ble | NcoI-SP-SalI,NotI-His6-stop | SalI-NotI | Signal Peptide | P14 | A26 | Amp/Ble/Tet |
|
|
V32 | pLEXSY-Ble | NcoI-SP-Strep Tag-His8-TEV-SalI,NotI-His6-stop | SalI-NotI | SP-Strep Tag-His8-TEV | P14 | A26 | Amp/Ble/Tet |
|
|
V33 | pLEXSY-Ble | NcoI-SalI,NotI-His8-stop | SalI-NotI | none | P14 | A26 | Amp/Ble/Tet |
|
|
V34 | pLEXSY-Ble | NcoI-Strep Tag-SalI,NotI-His8-stop | SalI-NotI | Strep Tag | P14 | A26 | Amp/Ble/Tet |
|
|
V35 | pLEXSY-Ble | NcoI-His8-GST-TEV-SalI,NotI-stop | SalI-NotI | His8-GST-TEV | P14 | A26 | Amp/Ble/Tet |
|
|
V36 | pLEXSY-Ble | NcoI-SP-Strep-His8-TEV-SalI,NotI-His6-stop | SalI-NotI | SP-Strep-His8-TEV | P14 | A26 | Amp/Ble/Tet |
|
|
V37 | pLEXSY-Ble | NcoI-Strep-His8-TEV-SalI,NotI-His6-stop | SalI-NotI | Strep-His8-TEV | P14 | A26 | Amp/Ble/Tet |
|
|
V40 | pTT5 | NcoI-His8-GB1-TEV-SalI,NotI-stop | SalI-NotI | His8-GB1-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V41 | pTT5 | NcoI-His6-Strep-His8-TEV-SalI,NotI-stop | SalI-NotI | His6-Strep-His8-TEV | P14 | A26 | Amp/Ble/Tet |
|
|
V42 | pTT5 | NcoI-Strep-His8-TEV-SalI,NotI-stop | SalI-NotI | Strep-His8-TEV | P14 | A26 | Amp/Ble/Tet |
|
|
V51 | pFab | EcoRV-SP-Strep-His8-TEV-SalI,NotI-His6-stop | SalI-NotI | SP-Strep-His8-TEV | unknown | unknown | Amp |
|
|
V52 | pFab | EcoRV-SP-His6-TEV-SalI,NotI-stop | SalI-NotI | SP-His6-TEV | unknown | unknown | Amp |
|
|
V53 | pFab | EcoRV-SP-His6-TEV-SalI,NotI-stop | SalI-NotI | SP-His6-TEV | unknown | unknown | Amp |
|
|
V54 | pFab | EcoRV-SP-His6-TEV-SalI,NotI-Strep-stop | SalI-NotI | SP-His6-TEV | unknown | unknown | Amp |
|
|
V55 | pMAL | NdeI-MBP-TEV-SalI,NotI-stop | SalI-NotI | MBP-TEV | M13R-pUC | M13F-pUC | Amp |
|
|
V56 | pMW103 | LexA-SalI,NotI-stop | SalI-NotI | LexA | unknown | unknown | Kan | For Y2H |
|
V57 | pET30a | NdeI-hDim2-SalI,NotI-XhoI-His6-stop | SalI-NotI | hDim2 | T7 Promoter | T7 Terminator | Kan |
|
|
V59 | pET30a | NdeI-His8-TEV-SalI,NotI-Lysozyme-stop | SalI-NotI | His8-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V60 | pET30a | NdeI-Lysozyme-SalI,NotI-XhoI-His6-stop | SalI-NotI | Lysozyme | T7 Promoter | T7 Terminator | Kan |
|
|
V61 | pEU-01 | His6-TEV-SalI,NotI-stop | SalI-NotI | His6-TEV | unknown | unknown | Amp |
|
|
V62 | pET30a | NdeI-MBP-SalI,NotI-lysozyme-XhoI-His6-stop | SalI-NotI | MBP | T7 Promoter | T7 Terminator | Kan |
|
|
V63 | pET30a | NdeI-MBP-TEV-SalI,NotI-Lysozyme-XhoI-His6-stop | SalI-NotI | MBP-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V64 | pET30a | NdeI-His8-MBP-TEV-SalI,NotI-Lysozyme-stop | SalI-NotI | His8-MBP-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V65 | pJG4-5 | Gal1-B42-HA epitope-EcoRI-SalI,NotI-stop | SalI-NotI | Gal1-B42-HA epitope | unknown | unknown | Amp | For Y2H |
|
V66 | pET30a | NdeI-T4LS5-SalI,NotI-XhoI-His6-stop | SalI-NotI | T4LS5 | T7 Promoter | T7 Terminator | Kan |
|
|
V67 | pEU-01 | NdeI-His8-NcoI-TEV-SalI,NotI-stop | SalI-NotI | His8-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V71 | pET30a | NdeI-SalI,NotI-XhoI-His6-stop | SalI-NotI | none | T7 Promoter | T7 Terminator | Kan |
|
|
V72 | pET30a | NdeI-SalI,NotI-Strep-stop | SalI-NotI | none | T7 Promoter | T7 Terminator | Kan |
|
|
V77 | pBAD | NdeI-SalI,NotI-XhoI-His6-stop | SalI-NotI | none | pBAD for | pBAD rev | Amp |
|
|
V78 | pBAD-V28E | NdeI-MBP-SalI,NotI-XhoI-His6-stop | SalI-NotI | MBP | pBAD for | pBAD rev | Amp |
|
|
V79 | pBAD-V29H | NdeI-His8-MBP-TEV-SalI,NotI-stop | SalI-NotI | His8-MBP-TEV | pBAD for | pBAD rev | Amp |
|
|
V80 | pCMV-HA | HAtag-EcoRI-SalI,NotI-stop | SalI-NotI | HAtag | pCMV | M13R-20 | Amp |
|
|
V81 | pBAD-V30 | NdeI-MBP-TEV-SalI,NotI-XhoI-His6-stop | SalI-NotI | MBP-TEV | pBAD for | pBAD rev | Amp |
|
|
V82 | pBAD-V29HS | NdeI-His8-MBP-TEV-SalI,NotI-strep-stop | SalI-NotI | His-MBP-TEV | pBAD for | pBAD rev | Amp |
|
|
V83 | pASK-17plus | NdeI-Strep-TEV-EcoRI-BamHI-XhoI-SalI-NcoI-EcoRV-HindIII | MCS | Strep-TEV | pASk for | pASK rev | Amp |
|
|
V84 | pASK-V3 | NdeI-His8-GB1-TEV-SalI,NotI-stop | SalI-NotI | His8-GB1-TEV | pASk for | pASK rev | Amp |
|
|
V85 | pASK-V28E | NdeI-MBP-SalI,NotI-XhoI-His6-stop | SalI-NotI | MBP | pASk for | pASK rev | Amp |
|
|
V86 | pASK-V29H | NdeI-His8-MBP-TEV-SalI,NotI-stop | SalI-NotI | His-MBP-TEV | pASk for | pASK rev | Amp |
|
|
V87 | pASK-V30 | NdeI-MBP-TEV-SalI,NotI-XhoI-His6-stop | SalI-NotI | MBP-TEV | pASk for | pASK rev | Amp |
|
|
V88 | pASK-V29HS | NdeI-His8-MBP-TEV-SalI,NotI-Strep-stop | SalI-NotI | His-MBP-TEV | pASk for | pASK rev | Amp |
|
|
V89 | pET30a | NdeI-InfB21-eGFP-SalI,NotI-stop | SalI-NotI | InfB21-eGFP | T7 Promoter | T7 Terminator | Kan |
|
|
V90 | pET30a | NdeI-InfB21-eGFP-TEV-SalI,NotI-stop | SalI-NotI | InfB21-eGFP | T7 Promoter | T7 Terminator | Kan |
|
|
V91 | pET30a | NdeI-His8-eGFP-TEV-SalI,NotI-stop | SalI-NotI | His8-eGFP-TEV | T7 Promoter | T7 Terminator | Kan |
|
|
V92 | pET30a | NdeI-SUMO-SalI,NotI-Strep-stop | SalI-NotI | SUMO | T7 Promoter | T7 Terminator | Kan |
|
|
V93 | pET30a | NdeI-His8-MBP-TEV-GFP-SalI,NotI-Strep-stop | SalI-NotI | His8-MBP-TEV-GFP | T7 Promoter | T7 Terminator | Kan |
|
|
V94 | pET30a | NdeI-VHb-SalI,NotI-Strep-stop | SalI-NotI | VHb | T7 Promoter | T7 Terminator | Kan |
|
|
V95 | pET30a | NdeI-Tag2-SalI,NotI-Strep-stop | SalI-NotI | Tag2 | T7 Promoter | T7 Terminator | Kan |
|
|
V96 | pET30a | NdeI-MBP-SalI,NotI-XhoI-His6-stop | SalI-NotI | MBP | T7 Promoter | T7 Terminator | Kan |
|
|
V97 | pET30a | NdeI-MBP-SalI,NotI-Strep-stop | SalI-NotI | MBP | T7 Promoter | T7 Terminator | Kan |
|
|
V98 | pFab | EcoRV-SP-NdeI-His8-SalI,NotI-Strep-stop | SalI-NotI | SP-His8 | unknown | unknown | Amp |
|
|
V99 | pET30a | NdeI-MBP-SalI,NotI-XhoI-His6-stop | SalI-NotI | MBP | T7 Promoter | T7 Terminator | Kan |
|
|
V100 | pET30a | NdeI-MBP-SalI,NotI-XhoI-His6-stop | SalI-NotI | MBP | T7 Promoter | T7 Terminator | Kan |
|
|
V101 | pET30a | NdeI-GlyA-SalI,NotI-XhoI-His6-stop | SalI-NotI | GlyA | T7 Promoter | T7 Terminator | Kan |
|
|
V102 | pET30a | NdeI-GacH-SalI,NotI-XhoI-His6-stop | SalI-NotI | GacH | T7 Promoter | T7 Terminator | Kan |
|
|
V103 | pET30a | NdeI-XBP1-SalI,NotI-XhoI-His6-stop | SalI-NotI | XBP1 | T7 Promoter | T7 Terminator | Kan |
|
|
V104 | pGBKT7SN | DNA Binding-XhoI-NdeI-SalI,NotI-stop | SalI-NotI | DNA Binding | T7 Promoter | ADH | Kan |
|
|
V105 | pGADT7SN | SV40-AD-NcoI-Epitope-NdeI-SalI,NotI-stop | SalI-NotI | SV40-AD-Epitope | T7 Promoter | ADH | Amp |
|
|
V106 | pRmHa3 | EcoRI-TEV-SalI,NotI-XhoI-His6-stop | SalI-NotI | TEV | pRmHa3 for | pRmHa3 rev | Amp |
|
|
V107 | pVL1392 | NcoI-NotI-HIs6-stop | unknown | none | y326 polyhedrin | y327 polyhedrin | Amp |
|
|
V108 | plSUMOstar | HIs6-SUMOstar-SalI,NotI-stop | SalI-NotI | SUMOstar | unknown | unknown | Amp/Get |
|
|
V109 | pFastBacHT | NdeI-InfB21-His8-TEV-SalI,NotI-stop | SalI-NotI | InfB21-His8-TEV | pASk for | pASK rev | Amp/Get |
|
|
V110 | pFastBacHT | NdeI-SalI,NotI-XhoI-His6-stop | SalI-NotI | none | pASk for | pASK rev | Amp/Get |
|
|
V111 | pFastBacHT | NdeI-His8-MBP-TEV-Sali,NotI-stop | SalI-NotI | His8-MBP-TEV | pASk for | pASK rev | Amp/Get |
|
|
V112 | pFastBacHT | NdeI-His8-MBP-TEV-Sali,NotI-XhoI-His6-stop | SalI-NotI | His8-MBP-TEV | pASk for | pASK rev | Amp/Get |
|
|
V113 | pColdII | NdeI-His8-MBP-TEV-SalI,NotI-stop | SalI-NotI | His8-MBP-TEV | pCold for | pCold rev | Amp |
|
|
V114 | pColdII | NdeI-MBP-TEV-SalI,NotI-stop | SalI-NotI | MBP-TEV | pCold for | pCold rev | Amp |
|
|
V115 | pColdII | NdeI-XhoI-His6-stop | unknown | none | pCold for | pCold rev | Amp |
|
|
V116 | pET30a | NdeI-B562-XhoI-His6-stop | unknown | B562 | T7 Promoter | T7 Terminator | Kan |
|
|
V117 | pCold |
|
|
|
|
|
|
|
|
V118 | pGADzaA |
|
|
|
|
|
|
|
|
V119 | FBQH | His8-TEV-BamHI-XhoI | BamHI-XhoI | His8-TEV | FBQ-F | FBQ-R | Amp |
|
|
V120 | FBQH | GST-TEV-BamHI-Xho I | BamHI-XhoI | GST-TEV | FBQ-F | FBQ-R | Amp |
|
|
V121 | FBQH | His-MBP-TEV-BamHI-XhoI | BamHI-XhoI | His-MBP-TEV | FBQ-F | FBQ-R | Amp |
|
|
V122 | PET30a | NdeI- His8- GB1 -Sal I -Not I-stop | Sal I-Not I | His8- GB1 | T7-P | T7-T | Kan | XY |
|
V123 | pCDNA3.1 | CMV-T7-FLAG-MCS-Stop | XhoI-BamH I-EcoR I-Not I | FLAG | T7/CMV | BGH-R | Amp | XY |
|
V124 | pCDNA3.1 | CMV-T7-HA-MCS-Stop | XhoI-BamH I-EcoR I-Not I | HA | T7/CMV | BGH-R | Amp | XY |
|
V125 | pCDNA3.1 | CMV-T7-Strep-MCS-Stop | XhoI-BamH I-EcoR I-Not I | Strep | T7/CMV | BGH-R | Amp | XY |
|
V126 | pCDNA3.1 | CMV-T7-SP-3XFLAG-MCS-Stop | XhoI-BamH I-EcoR I-Not I | 3XFLAG | T7/CMV | BGH-R | Amp | XY | SP=signal peptide |
V127 | pCDNA3.1 | CMV-T7-SP-FLAG-Clover-MCS-Stop | XhoI-BamH I-EcoR I-Not I | Clover/FLAG | T7/CMV | BGH-R | Amp | XY | SP=signal peptide |
V128 | pCDNA3.1 | CMV-T7-SP-HA-MCS-Stop | XhoI-BamH I-EcoR I-Not I | HA | T7/CMV | BGH-R | Amp | XY | SP=signal peptide |
V129 | pCDNA3.1 | CMV-T7-SP-Strep-mRubby2-MCS-Stop | XhoI-BamH I-EcoR I-Not I | Strep/mRubby2 | T7/CMV | BGH-R | Amp | XY | SP=signal peptide |
V130 | pCDNA3.1 | CMV-T7-SP-MCS-Stop | XhoI-BamH I-EcoR I-Not I | Strep | T7/CMV | BGH-R | Amp | XY |
|
V131 | pCDNA3.1 | CMV-T7-Strep-mRubby2-MCS-Stop | XhoI-BamH I-EcoR I-Not I | Strep/mRubby2 | T7/CMV | BGH-R | Amp | XY |
|
V132 | pCDNA3.1 | CMV-T7-FLAG-Clover-MCS-Stop | XhoI-BamH I-EcoR I-Not I | Clover/FLAG | T7/CMV | BGH-R | Amp | XY |
|
V133 | pCDNA3.1 | CMV-T7-Strep-NEGFP-MCS-Stop | XhoI-Not I | Strep/NEGFP | T7/CMV | BGH-R | Amp | XY |
|
V134 | pCDNA3.1 | CMV-T7-MCS-CEGFP-Myc-Stop | XhoI-Not I | Myc/CEGFP | T7/CMV | BGH-R | Amp | XY |
|
V135 | PRSF | T7-His6-MCS-Stop/T7-MBP-TEV-MCS-His6-Stop | BamH I-Not I/Hind III-EcoR I | His/MBP | Z21 | Z20 | Kan | XY |
|
V136 | plsecSUMOstar | GP67A-HIs6-SUMOstar-SalI,NotI-stop | Sal I-Not I | His | Y935 | FXJ57 | Amp |
|
|
V137 | plsecSUMOstar | GP67A-HIs6-SUMOstar-MBP-TEV-SalI,NotI-stop | Sal I-Not I | His/MBP | Z8 | FXJ57 | Amp |
|
|
V138 | plsecSUMOstar | GP67A-HIs6-TEV-SalI,NotI-stop | Sal I-Not I | His-TEV | Z8 | FXJ57 | Amp |
|
|
V139 | plsecSUMOstar | GP67A-HIs6-SalI,NotI-stop | Sal I-Not I | His | FXJ66 |
| Amp |
|
|
V140 | PTT5 | IFNA1-SalI,XbaI-His6-stop | Sal I-Xba I | His | PTT5-F | PTT5-R | Amp |
|
|
V141 | PTT5 | IFNA1-SalI,XbaI-Fc-stop | Sal I-Xba I | Fc | PTT5-F | PTT5-R | Amp |
|
|
V142 | PTT5 | IFNA1-SalI,XbaI-Fc-TEV-stop | Sal I-Xba I | Fc-TEV | PTT5-F | PTT5-R | Amp |
|
|
V143 | plsecSUMOstar | GP67A-HIs6-SUMOstar-EGFP-TEV-SalI,NotI-stop | Sal I-Not I | His-EGFP-TEV | Z8 | FXJ57 | Amp |
|
|
1. DH5α菌株
基因型:F-、φ80dlacZΔM15、Δ(lacZYA-argF)U169、deoR、recA1、endA1、hsdR17(rk-,
mk+)、phoA、supE44、λ-、thi-1、gyrA96、relA1。
DH5α是一种常用于质粒克隆的菌株。E.coli DH5α在使用pUC系列质粒载体转化时,其Φ80dlacZΔM15基因的表达产物与pUC载体编码的β-半乳糖苷酶氨基端实现α互补,可用于蓝白斑筛选。recA1和endA1的突变有利于克隆DNA的稳定和高纯度质粒DNA的提取。用于分子克隆、质粒提取和蛋白质表达。
2. TOP10菌株
基因型:F- 、mcrAΔ(mrr-hsd RMS-mcrBC)、 φ80,lacZΔM15、△lacⅩ74、 recA1,araΔ139Δ(ara-leu)7697、galU、galK、rps、(Strr) endA1、nupG。
该菌株适用于高效的DNA克隆和质粒扩增,能保证高拷贝质粒的稳定遗传。
3. HB101菌株
基因型:supE44、hsdS20(rB-mB-)、recA13、ara-14、proA2、lacY1、galK2、rpsL20、xyl-5、mtl-1、leuB6,thi-1。该菌株遗传性能稳定,使用方便,适用于各种基因重组实验。
4. XL1-Blue菌株
基因型:endA1、gyrA96(nalR)、thi-1、recA1、relA1、lac、glnV44、F′[Tn10 proAB+ lacIq Δ(lacZ)M15]、hsdR17(rK- mK+)。该菌株具有卡那抗性、四环素抗性和氯霉素抗性,用于分子克隆和质粒提取。
5. DH10B菌株
基因型:F-、mcrA、Δ(mrr-hsdRMS-mcrBC)、Φ80dlacZΔM15、ΔlacX74、endA1、recA1、deoR、Δ(ara,leu)7697 araD139、galU、galK、nupG、rpsL、λ-。
该菌株消除了mcrA、mcrBC、mrr和hsdRMS限制系统,允许构建更多具有代表性的基因组文库。endA1突变,可以增加质粒产量和数量。高效率转化大小为150 kb的质粒,用于产生cDNA或基因组文库。lacZΔM15 用于重组克隆的蓝/白斑筛选。
6. B834菌株
是BL21菌株的父代菌株。适用于用Se-Met方法表达硒代蛋白。无抗性。
7. BL21(DE3)菌株
基因型:F-、ompT、hsdS(rBB-mB-)、gal、dcm(DE3)
该菌株用于高效表达克隆于含有噬菌体T7启动子的表达载体(如pET系列)的基因。T7噬菌体RNA聚合酶位于λ 噬菌体DE3区,该区整合于BL21的染色体上。该菌适合表达非毒性蛋白。
8. BL21(DE3) pLysS菌株
基因型:F-、ompT hsdS(rBB-mB-)、gal、dcm(DE3)、pLysS、Camr
该菌株含有质粒pLysS,因此具有氯霉素抗性。PLysS含有表达T7溶菌酶的基因,能够通过直接与T7 RNA 聚合酶结合并抑制其活性来降低目的基因的背景表达水平,但不干扰IPTG诱导的表达。该菌适合表达毒性蛋白和非毒性蛋白。需要用更高浓度IPTG比如0.7 mM来诱导。
9. BL21(DE3) RosettaTM菌株(Novagen)
Genotype:F-、ompT、hsdSB(rB- mB-)、gal、dcm (DE3)、pRARE (CamR)。该菌株是BL21额外含有一个编码稀有密码子的质粒,因此具有氯霉素抗性。
Rosetta™ host strains are BL21 derivatives designed to enhance the expression of eukaryotic proteins that contain codons rarely used in E. coli. These strains supply tRNAs for AGG, AGA, AUA, CUA, CCC, GGA codons on a compatible chloramphenicol-resistant plasmid. Thus the Rosetta strains provide for “universal” translation which is otherwise limited by the codon usage of E. coli. The tRNA genes are driven by their native promoters.
10. BL21 Codon plusTM(DE3)-RIPL (Stratagene)
Antibiotic Resistance:Camr Strep/Specr 该菌株是BL21额外含有一个编码稀有密码子的质粒,因此具有氯霉素抗性。
BL21-CodonPlus competent cells are derived from the high-performance Stratagene BL21-Gold competent cell line. These cells enable efficient high-level expression of heterologous proteins in Escherichia coli.
BL21-CodonPlus strains are engineered to contain extra copies of genes that encode the tRNAs that most frequently limit translation of heterologous proteins in E. coli. Availability of tRNAs allows high-level expression of many heterologous recombinant genes in BL21-CodonPlus cells that are poorly expressed in conventional BL21 strains. The BL21-CodonPlus (DE3)-RIPL cells contain extra copies of the argU, ileY, and leuW as well as the + tRNA genes. This strain rescues expression of heterologous proteins from organisms that have either AT- or GC-rich genomes
11. JM109菌株
基因型:recA1、endA1、gyrA96、thi-1、hsdR17、supE44、relA1、Δ(lac-proAB)/F’[traD36、proAB+、lacIq、lacZΔM15]。该菌株在使用pUC系列质粒载体进行DNA转化或用M13 phage载体进行转染时,由于载体DNA产生的LacZa多肽和JM09编码的LacZΔM15进行α-互补,从而显示β-半乳糖苷酶活性,由此很容易鉴别重组体菌株。部分抗性缺陷,适合重复基因表达, 可用于M13克隆序列测定和蓝白斑筛选。用于分子克隆、质粒提取和蛋白质表达。
12. 293T细胞
293T细胞来源都是人胚胎肾细胞,通过基因技术派生的细胞系,经过腺病毒E1A基因转染,能表达SV40大T抗原,含有SV40复制起始点与启动子区。许多真核表达载体如pcDNA3.1中含有SV40病毒的复制起始位点,可在表达SV40病毒T抗原的细胞系中复制,从而提高外源基因的表达水平。因此293T细胞广泛用于病毒包装。用磷酸钙转染效率可高达50%。蛋白表达水平高,转染后2-3天用碱性磷酸酶分析可较容易地检测到表达的蛋白。瞬转293T细胞是过表达蛋白并获得细胞内及细胞外(分泌的或膜)蛋白的便捷方式。
Table of Cell Strains
Table :Cell strains
| Cell Strains | Resistance | Usage |
1 | Sf9 | none | 包装病毒 |
2 | Hi5 | none | 昆虫细胞表达蛋白 |
3 | HEK293 | none | 表达包装病毒 |
4 | 293T | none | 工具细胞 |
5 | L929 | none | 分泌gmcsf,促进巨噬细胞分化 |
Table of Bacteria Strains
Table :Bacteria Strains
| Bacteria Strains | Resistance | Usage |
1 | DH5α | none | Plasmid transformation |
2 | Top 10 | none | Plasmid transformation |
3 | TOP 10 F’ | Tet | Plasmid transformation |
4 | XL1blue MRF’ Kan | Kan | Plasmid transformation |
5 | xl-10 gold | Tet & Chl | Plasmid transformation |
6 | ccdB survival 2 T1R, Invitrogen | Sm | Plasmid transformation |
7 | Survival-2 | Sm | Plasmid transformation |
8 | Ex0-DH5α Amp | No found | Plasmid transformation |
9 | Lig-DH5α Amp | No found | Plasmid transformation |
10 | BL21(DE3) | none | Protein expression |
11 | BL21(DE3), pLysS | Chl | Protein expression |
12 | BL21* | none | Protein expression |
13 | BL21/AI Invitrogen | Tet | Protein expression |
15 | BL21,RIL,RIPL | Chl | Protein expression |
16 | Origami B | Kan & Tet | Protein expression |
17 | Rosetta Gami | Kan & Tet & Chl | Protein expression |
18 | Tuner | none | Protein expression |
19 | B834 | none | Protein expression |
20 | 55244 | Kan | Protein expression |
21 | Tuvp20 | No found |
|
22 | AH109 (Yeast) | none | 酿酒酵母,酵母双杂交,蛋白与蛋白互作 |
23 | SMD1168H (Yeast) | none | 毕赤酵母,蛋白表达 |
24 | CG1945 (Yeast) | none | 酿酒酵母,酵母双杂交,蛋白与蛋白互作,需要mating |
▲ 表格中”No found” 部分因未查到其细胞抗性,建议实验中视为无抗性处理,其具体特性请自行查询。
1.2.8 结晶试剂盒
简介(INTRODUCTION)
长晶体是由清澈的蛋白质溶液变成饱和溶液,饱和溶液变成过饱和溶液,过饱和溶液如果满足“适当的”条件时蛋白质有可能以晶体形式从溶液中析出,否则就会变成无定形沉淀。而结晶过程不可能自发地完成,必须要在外界的“力”帮助下才可能完成这种熵减的过程。
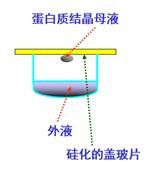 |
我们实验室用的是悬滴气象扩散法,在起始状态时,结晶母液与外液之间存在某些“因素”的梯度差,随着气相扩散,这些梯度发生变化。如果变化后的条件合适,母液中的蛋白质就有可能以晶体状态析出。
我们实验室有很多长晶体试剂盒,还有一个金老师自己研制出来的MBP-screen。原则上带MBP标签的蛋白一开始可以用MBP-screen来筛选,如果没有晶体长出来时,根据蛋白在该试剂盒的表现(都是沉淀还是都是清澈的液滴)来决定下一步用什么试剂盒。本附录中收纳了我们实验室已有的所有试剂盒的配方。

实验室现有晶体试剂盒介绍
1. MBP-phusion Protein Crystallization Screen
本实验室研制,利用率和成功率最高的试剂盒。是MBP偶联蛋白的首选,其他样品也适用,一般用于初筛。

2. Wizard™ I/Ⅱrandom sparse matrix crystallization screen
3. Wizard™ III/Ⅳrandom sparse matrix crystallization screen
适合初筛,pH值横跨4.5-10.5,不同的buffer,不同的盐,沉淀剂也有很多种。

4. The Anions Suite
该试剂盒组分都是盐离子,没有以PEG,EG或者甘油作为沉淀剂。如果蛋白在初筛的过程中,以PEG为
沉淀剂的情况下极其容易沉淀时,可以用该试剂盒试一下。
5. The Nucleix Suite
相对来说是核酸-蛋白复合物的首选试剂盒。
6. The PACT Suite
以不同的PEG为沉淀剂的试剂盒,如果初筛后发现你的蛋白在以PEG为沉淀剂(PEG 1000以上)的情况下长晶体的可能性较大时,这个试剂盒可以帮你确定哪个PEG搭配哪个盐离子比较好。
7. The Protein Complex Suite
顾名思义,适合蛋白复合物的晶体筛选,但同时也适合来自哺乳动物的蛋白。

8. Natrix HT(HR2-131)
主要用于核酸以及核酸-蛋白质复合物
9. SaltRx HT(HR2-136)
该试剂盒与Qiagen的Anions试剂盒相似,主要成分全部都是盐离子,其中包含22中盐离子,配与不同的浓度和pH环境,在初筛中发现蛋白倾向于在盐离子环境下长晶体时可以考虑这个试剂盒。
10. PEGRx HT(HR2-086)
用于生物大分子的晶体筛选,其实也是比较适合用于初筛。
11. Crystal Screen HT(HR2-130)
该试剂盒是非常经典的一款试剂盒,里面包含不同的沉淀剂,不同的盐离子以及不同的pH环境,非常适合初筛。
Wizard™ I random sparse matrix crystallization screen
No. | crystallant | salt (0.2 M) | buffer (0.1 M) |
1 | 20% (w/v) PEG-8000 | none | CHES pH 9.5 |
2 | 10% (v/v) 2-propanol | NaCl | HEPES pH 7.5 |
3 | 15% (v/v) ethanol | none | CHES pH 9.5 |
4 | 35% (v/v) 2-methyl-2,4-pentanediol | MgCl2 | imidazole pH 8.0 |
5 | 30% (v/v) PEG-400 | none | CAPS pH 10.5 |
6 | 20% (w/v) PEG-3000 | none | citrate pH 5.5 |
7 | 10% (w/v) PEG-8000 | Zn(OAc)2 | MES pH 6.0 |
8 | 2.0 M (NH4)2SO4 | none | citrate pH 5.5 |
9 | 1.0 M (NH4)2HPO4 | none | acetate pH 4.5 |
10 | 20% (w/v) PEG-2000 MME | none | Tris pH 7.0 |
11 | 20% (v/v) 1,4-butanediol | Li2SO4 | MES pH 6.0 |
12 | 20% (w/v) PEG-1000 | Ca(OAc)2 | imidazole pH 8.0 |
13 | 1.26 M (NH4)2SO4 | none | cacodylate pH 6.5 |
14 | 1.0 M sodium citrate | none | cacodylate pH 6.5 |
15 | 10% (w/v) PEG-3000 | Li2SO4 | imidazole pH 8.0 |
16 | 2.5 M NaCl | none | Na/K phosphate pH 6.2 |
17 | 30% (w/v) PEG-8000 | Li2SO4 | acetate pH 4.5 |
18 | 1.0 M K/Na tartrate | NaCl | imidazole pH 8.0 |
19 | 20% (w/v) PEG-1000 | none | Tris pH 7.0 |
20 | 0.4 M NaH2PO4/1.6 M K2HPO4 | NaCl | imidazole pH 8.0 |
21 | 20% (w/v) PEG-8000 | none | HEPES pH 7.5 |
22 | 10% (v/v) 2-propanol | none | Tris pH 8.5 |
23 | 15% (v/v) ethanol | MgCl2 | imidazole pH 8.0 |
24 | 35% (v/v) 2-methyl-2,4-pentanediol | NaCl | Tris pH 7.0 |
25 | 30% (v/v) PEG-400 | MgCl2 | Tris pH 8.5 |
26 | 10% (w/v) PEG-3000 | none | CHES pH 9.5 |
27 | 1.2 M NaH2PO4/0.8 M K2HPO4 | Li2SO4 | CAPS pH 10.5 |
28 | 20% (w/v) PEG-3000 | NaCl | HEPES pH 7.5 |
29 | 10% (w/v) PEG-8000 | NaCl | CHES pH 9.5 |
30 | 1.26 M (NH4)2SO4 | NaCl | acetate pH 4.5 |
31 | 20% (w/v) PEG-8000 | NaCl | phosphate-citrate pH 4.2 |
32 | 10% (w/v) PEG-3000 | none | Na/K phosphate pH 6.2 |
33 | 2.0 M (NH4)2SO4 | Li2SO4 | CAPS pH 10.5 |
34 | 1.0 M (NH4)2HPO4 | none | imidazole pH 8.0 |
35 | 20% (v/v) 1,4-butanediol | none | acetate pH 4.5 |
36 | 1.0 M sodium citrate | none | imidazole pH 8.0 |
37 | 2.5 M NaCl | none | imidazole pH 8.0 |
38 | 1.0 M K/Na tartrate | Li2SO4 | CHES pH 9.5 |
39 | 20% (w/v) PEG-1000 | Li2SO4 | phosphate-citrate pH 4.2 |
40 | 10% (v/v) 2-propanol | Ca(OAc)2 | MES pH 6.0 |
41 | 30% (w/v) PEG-3000 | none | CHES pH 9.5 |
42 | 15% (v/v) ethanol | none | Tris pH 7.0 |
43 | 35% (v/v) 2-methyl-2,4-pentanediol | none | Na/K phosphate pH 6.2 |
44 | 30% (v/v) PEG-400 | Ca(OAc)2 | acetate pH 4.5 |
45 | 20% (w/v) PEG-3000 | none | acetate pH 4.5 |
46 | 10% (w/v) PEG-8000 | Ca(OAc)2 | imidazole pH 8.0 |
47 | 1.26 M (NH4)2SO4 | Li2SO4 | Tris pH 8.5 |
48 | 20% (w/v) PEG-1000 | Zn(OAc)2 | acetate pH 4.5 |
Wizard™ I random sparse matrix crystallization screen
No. | crystallant | salt (0.2 M) | buffer (0.1 M) |
49 | 10% (w/v) PEG-3000 | Zn(OAc)2 | acetate pH 4.5 |
50 | 35% (v/v) 2-methyl-2,4-pentanediol | Li2SO4 | MES pH 6.0 |
51 | 20% (w/v) PEG-8000 | MgCl2 | Tris pH 8.5 |
52 | 2.0 M (NH4)2SO4 | NaCl | cacodylate pH 6.5 |
53 | 20% (v/v) 1,4-butanediol | NaCl | HEPES pH 7.5 |
54 | 10% (v/v) 2-propanol | Li2SO4 | phosphate-citrate pH 4.2 |
55 | 30% (w/v) PEG-3000 | NaCl | Tris pH 7.0 |
56 | 10% (w/v) PEG-8000 | NaCl | Na/K phosphate pH 6.2 |
57 | 2.0 M (NH4)2SO4 | none | phosphate-citrate pH 4.2 |
58 | 1.0 M (NH4)2HPO4 | none | Tris pH 8.5 |
59 | 10% (v/v) 2-propanol | Zn(OAc)2 | cacodylate pH 6.5 |
60 | 30% (v/v) PEG-400 | Li2SO4 | cacodylate pH 6.5 |
61 | 15% (v/v) ethanol | Li2SO4 | citrate pH 5.5 |
62 | 20% (w/v) PEG-1000 | NaCl | Na/K phosphate pH 6.2 |
63 | 1.26 M (NH4)2SO4 | none | HEPES pH 7.5 |
64 | 1.0 M sodium citrate | none | CHES pH 9.5 |
65 | 2.5 M NaCl | MgCl2 | Tris pH 7.0 |
66 | 20% (w/v) PEG-3000 | Ca(OAc)2 | Tris pH 7.0 |
67 | 1.6 M NaH2PO4/0.4 M K2HPO4 | none | phosphate-citrate pH 4.2 |
68 | 15% (v/v) ethanol | Zn(OAc)2 | MES pH 6.0 |
69 | 35% (v/v) 2-methyl-2,4-pentanediol | none | acetate pH 4.5 |
70 | 10% (v/v) 2-propanol | none | imidazole pH 8.0 |
71 | 15% (v/v) ethanol | MgCl2 | HEPES pH 7.5 |
72 | 30% (w/v) PEG-8000 | NaCl | imidazole pH 8.0 |
73 | 35% (v/v) 2-methyl-2,4-pentanediol | NaCl | HEPES pH 7.5 |
74 | 30% (v/v) PEG-400 | none | CHES pH 9.5 |
75 | 10% (w/v) PEG-3000 | MgCl2 | cacodylate pH 6.5 |
76 | 20% (w/v) PEG-8000 | Ca(OAc)2 | MES pH 6.0 |
77 | 1.26 M (NH4)2SO4 | NaCl | CHES pH 9.5 |
78 | 20% (v/v) 1,4-butanediol | Zn(OAc)2 | imidazole pH 8.0 |
79 | 1.0 M sodium citrate | NaCl | Tris pH 7.0 |
80 | 20% (w/v) PEG-1000 | none | Tris pH 8.5 |
81 | 1.0 M (NH4)2HPO4 | NaCl | citrate pH 5.5 |
82 | 10% (w/v) PEG-8000 | none | imidazole pH 8.0 |
83 | 0.8 M NaH2PO4/1.2 M K2HPO4 | none | acetate pH 4.5 |
84 | 10% (w/v) PEG-3000 | NaCl | phosphate-citrate pH 4.2 |
85 | 1.0 M K/Na tartrate | Li2SO4 | Tris pH 7.0 |
86 | 2.5 M NaCl | Li2SO4 | acetate pH 4.5 |
87 | 20% (w/v) PEG-8000 | NaCl | CAPS pH 10.5 |
88 | 20% (w/v) PEG-3000 | Zn(OAc)2 | imidazole pH 8.0 |
89 | 2.0 M (NH4)2SO4 | Li2SO4 | Tris pH 7.0 |
90 | 30% (v/v) PEG-400 | NaCl | HEPES pH 7.5 |
91 | 10% (w/v) PEG-8000 | MgCl2 | Tris pH 7.0 |
92 | 20% (w/v) PEG-1000 | MgCl2 | cacodylate pH 6.5 |
93 | 1.26 M (NH4)2SO4 | none | MES pH 6.0 |
94 | 1.0 M (NH4)2HPO4 | NaCl | imidazole pH 8.0 |
95 | 2.5 M NaCl | Zn(OAc)2 | imidazole pH 8.0 |
96 | 1.0 M K/Na tartrate | none | MES pH 6.0 |
Wizard™ III random sparse matrix crystallization screen
No | crystallant | salt/additive #1 | additive #2 | buffer | pH |
1 | 20%(w/v) PEG 3350 | 0.2 M ammonium citrate (dibasic) |
|
|
|
2 | 30%(v/v) MPD | 0.02 M calcium chloride |
| 0.1 M sodium acetate | 4.6 |
3 | 20%(w/v) PEG 3350 | 0.2 M magnesium formate |
|
|
|
4 | 20%(w/v) PEG 3350 | 0.2 M ammonium formate |
|
|
|
5 | 20%(w/v) PEG 3350 | 0.2 M ammonium chloride |
|
|
|
6 | 20%(w/v) PEG 3350 | 0.2 M potassium formate |
|
|
|
7 | 50%(w/v) MPD | 0.2 M NH4H2PO4 |
| 0.1 M Tris | 8 |
8 | 20%(w/v) PEG 3350 | 0.2 M potassium nitrate | |||
9 | 0.8 M ammonium sulfate | 0.1 M citric acid | 4.0 | ||
10 | 20%(w/v) PEG 3350 | 0.2 M sodium thiocyanate | |||
11 | 20%(w/v) PEG 6000 | 0.1 M bicine | 9.0 | ||
12 | 10%(w/v) PEG 8000 | 8%(v/v) ethylene glycol | 0.1 M HEPES | 7.5 | |
13 | 8%(w/v) PEG 4000 | 0.1 M sodium acetate | 4.6 | ||
14 | 20%(w/v) PEG 6000 | 0.1 M citric acid | 5.0 | ||
15 | 1.6 M Sodium citrate | ||||
16 | 20%(w/v) PEG 3350 | 0.2 M potassium citrate tribasic | |||
17 | 20%(w/v) PEG 4000 | 10%(v/v) 2-propanol | 0.1 M citrate | 5.5 | |
18 | 20%(w/v) PEG 6000 | 1 M lithium chloride | 0.1 M citric acid | 4.0 | |
19 | 20%(w/v) PEG 3350 | 0.2 M ammonium nitrate | |||
20 | 10%(w/v) PEG 6000 | 0.1 M HEPES | 7.0 | ||
21 | 1.6 M Na/K phosphate | 0.1 M HEPES | 7.5 | ||
22 | 20%(v/v) ethanol | 0.1 M Tris | 8.5 | ||
23 | 10%(w/v) PEG 20,000 | 2%(v/v) dioxane | 0.1 M bicine | 9.0 | |
24 | 2 M ammonium sulfate | 0.1 M sodium acetate | 4.6 | ||
25 | 10%(w/v) PEG 1000 | 10%(w/v) PEG 8000 | |||
26 | 24%(w/v) PEG 1500 | 20%(v/v) glycerol | |||
27 | 30%(v/v) PEG 400 | 0.2 M magnesium chloride | 0.1 M HEPES | 7.5 | |
28 | 70%(v/v) MPD | 0.1 M HEPES | 7.5 | ||
29 | 40%(v/v) MPD | 0.1 M Tris | 8.0 | ||
30 | 5.5%(w/v) PEG 4000 | 0.17 M ammonium sulfate | 15%(v/v) glycerol | ||
31 | 14%(v/v) 2-propanol | 0.14 M calcium chloride | 30%(v/v) glycerol | 0.07 M sodium acetate | 4.6 |
32 | 16%(w/v) PEG 8000 | 0.04 M KH2PO4 | 20%(v/v) glycerol | ||
33 | 1.6 M magnesium sulfate | 0.1 M MES | 6.5 | ||
34 | 10%(w/v) PEG 6000 | 0.1 M bicine | 9.0 | ||
35 | 4.4%(w/v) PEG 8000 | 0.16 M calcium acetate | 20%(v/v) glycerol | 0.08 M cacodylate | 6.5 |
36 | 30%(v/v) Jeffamine M-600 | 0.05 M cesium chloride | 0.1 M MES | 6.5 | |
37 | 3.2 M ammonium sulfate | 0.1 M citric acid | 5.0 | ||
38 | 15%(w/v) PEG 10,000 | 2%(v/v) dioxane | 0.1 M citrate | 5.5 | |
39 | 20%(v/v) Jeffamine M-600 | 0.1 M HEPES | 7.5 | ||
40 | 10%(v/v) MPD | 0.1 M bicine | 9.0 | ||
41 | 28%(v/v) PEG 400 | 0.2 M calcium chloride | 0.1 M HEPES | 7.5 | |
42 | 30%(w/v) PEG 4000 | 0.2 M lithium sulfate | 0.1 M Tris | 8.5 | |
43 | 30%(w/v) PEG 8000 | 0.2 M ammonium sulfate | |||
44 | 30%(w/v) PEG 5000 MME | 0.2 M lithium sulfate | 0.1 M Tris | 8.0 | |
45 | 1.5 M ammonium sulfate | 12%(v/v) glycerol | 0.1 M Tris | 8.5 | |
46 | 50%(v/v) MPD | 0.2 M NH4H2PO4 |
| 0.1 M Tris | 8.5 |
47 | 30%(w/v) PEG 5000 MME | 0.2 M ammonium sulfate |
| 0.1 M MES | 6.5 |
48 | 20%(w/v) PEG 10,000 |
|
|
| 7.5 |
Wizard™ III random sparse matrix crystallization screen
No | crystallant | salt/additive #1 | additive #2 | buffer | pH |
49 | 20%(v/v) Glycerol | 40 mM Potassium phosphate | 16%(w/v) PEG 8000 | ||
50 | 15%(v/v) Ethanol | 100 mM Sodium chloride | 5%(v/v) MPD | Tris | 8.0 |
51 | 40%(v/v) Ethanol | 5%(w/v) PEG 1000 | Phosphate-citrate | 4.2 | |
52 | 200 mM Ammonium sulfate | BisTris | 5.5 | ||
53 | 2 M Ammonium sulfate | 2%(v/v) PEG 400 | Acetate | 5.5 | |
54 | 800 mM Ammonium Sulfate | Citrate | 4.0 | ||
55 | 2 M Lithium sulfate | 100 mM Magnesium sulfate | 5%(v/v) Isopropanol | Acetate | 4.5 |
56 | 2 M Lithium sulfate | 2%(v/v) PEG 400 | Tris | 8.5 | |
57 | 2 M Lithium sulfate | 100 mM Magnesium sulfate | 5%(v/v) PEG 400 | Acetate | 5.5 |
58 | 50%(v/v) PEG 200 | 200 mM Magnesium chloride | Na Cacodylate | 6.5 | |
59 | 40%(v/v) PEG 300 | 200 mM Calcium acetate | Na Cacodylate | 6.5 | |
60 | 30%(v/v) Jeffamine M600 pH 7.0 | 200 mM Lithium sulfate | HEPES | 7.0 | |
61 | 800 mM Succinic Acid pH 7.0 | ||||
62 | 40%(v/v) PEG 400 | 200 mM Lithium sulfate | Tris | 8.5 | |
63 | 50%(v/v) PEG 400 | 200 mM Lithium sulfate | Acetate | 4.5 | |
64 | 15%(v/v) PEG 550 MME | MES | 6.5 | ||
65 | 25%(w/v) PEG 1500 | SPG Buffer/NaOH | 5.5 | ||
66 | 25%(w/v) PEG 1500 | SPG Buffer/NaOH | 8.5 | ||
67 | 25%(w/v) PEG 1500 | MMT Buffer/NaOH | 6.5 | ||
68 | 25%(w/v) PEG 1500 | MMT Buffer/NaOH | 9.0 | ||
69 | 25%(w/v) PEG 1500 | MIB Buffer/HCl | 5.0 | ||
70 | 25%(w/v) PEG 1500 | PCB Buffer/NaOH | 7.0 | ||
71 | 12%(w/v) PEG 1500 | 2500 mM Sodium chloride | 1.5%(v/v) MPD | Na Acetate | 5.5 |
72 | 2400 mM Sodium Malonate | ||||
73 | 30%(w/v) PEG 2000 MME | 150 mM Potassium bromide | |||
74 | 10%(w/v) PEG 2000 MME | 200 mM Ammonium sulfate | Sodium acetate | 5.5 | |
75 | 20%(w/v) PEG 2000 MME | 200 mM Trimethylamine n-oxide | Tris | 8.5 | |
76 | 20%(w/v) PEG 3350 | 200 mM Sodium fluoride | BisTris Propane | 6.5 | |
77 | 20%(w/v) PEG 3350 | 200 mM Sodium citrate | Citrate | 4.0 | |
78 | 20%(w/v) PEG 3350 | 200 mM Sodium malonate | BisTris Propane | 8.5 | |
79 | 20%(w/v) Polyacrylic acid 5100 | 20 mM Magnesium chloride | HEPES | 7.0 | |
80 | 2100 mM DL Malic acid pH 7.0 | ||||
81 | 800 mM K2HPO4 | 800 mM Na3PO4 | HEPES | 7.5 | |
82 | 20%(w/v) PEG 6000 | 200 mM Ammonium chloride | MES | 6.0 | |
83 | 20%(w/v) PEG 6000 | 200 mM Sodium chloride | HEPES | 7.0 | |
84 | 20%(w/v) PEG 6000 | 200 mM Lithium chloride | Tris | 8.0 | |
85 | 20%(w/v) Polyvinylpyrrolidone K15 | 100 mM Cobalt chloride | Tris | 8.5 | |
86 | 50%(v/v) Ethylene glycol | 200 mM Magnesium chloride | Tris | 8.5 | |
87 | 20%(w/v) PEG 8000 | 3%(v/v) MPD | Imidazole | 6.5 | |
88 | 20%(w/v) PEG 8000 | 100 mM Magnesium chloride | 20%(v/v) PEG 400 | Tris | 8.5 |
89 | 20%(w/v) PEG 8000 | 200 mM Ammonium sulfate | 10%(v/v) Isopropanol | HEPES | 7.5 |
90 | 30%(v/v) MPD | 25%(w/v) PEG 1500 | Acetate | 4.5 | |
91 | 30%(v/v) MPD | 200 mM Ammonium sulfate | 10%(w/v) PEG 3350 | Imidazole | 6.5 |
92 | 30%(v/v) MPD | 500 mM Sodium chloride | 8%(w/v) PEG 8000 | Tris | 8.5 |
93 | 40%(v/v) Isopropanol | 15%(w/v) PEG 8000 | Imidazole-cl | 6.5 | |
94 | 30%(v/v) Isopropanol | 30%(w/v) PEG 3350 | Tris | 8.5 | |
95 | 17%(w/v) PEG 10000 | 100 mM Ammonium Acetate | BisTris | 5.5 | |
96 | 15%(w/v) PEG 20000 | HEPES | 7.0 | ||
The Anions Suite
No | Precipitant | Salt | Buffer |
1 | 2.5 M Sodium acetate | 0.1 M Sodium acetate pH 4.6 | |
2 | 1.25 M Sodium acetate | 0.1 M Sodium acetate pH 4.6 | |
3 | 1.2 M tri-Sodium citrate | 0.1 M Sodium acetate pH 4.6 | |
4 | 0.6 M tri-Sodium citrate | 0.1 M Sodium acetate pH 4.6 | |
5 | 0.6 M Sodium fluoride | 0.1 M Sodium acetate pH 4.6 | |
6 | 0.3 M Sodium fluoride | 0.1 M Sodium acetate pH 4.6 | |
7 | 2.5 M Sodium acetate | 0.1 M MES pH 6.5 | |
8 | 1.25 M Sodium acetate | 0.1 M MES pH 6.5 | |
9 | 1.2 M tri-Sodium citrate | 0.1 M MES pH 6.5 | |
10 | 0.6 M tri-Sodium citrate | 0.1 M MES pH 6.5 | |
11 | 0.6 M Sodium fluoride | 0.1 M MES pH 6.5 | |
12 | 0.3 M Sodium fluoride | 0.1 M MES pH 6.5 | |
13 | 2.5 M Sodium acetate | 0.1 M HEPES pH 7.5 | |
14 | 1.25 M Sodium acetate | 0.1 M HEPES pH 7.5 | |
15 | 1.2 M tri-Sodium citrate | 0.1 M HEPES pH 7.5 | |
16 | 0.6 M tri-Sodium citrate | 0.1 M HEPES pH 7.5 | |
17 | 0.6 M Sodium fluoride | 0.1 M HEPES pH 7.5 | |
18 | 0.3 M Sodium fluoride | 0.1 M HEPES pH 7.5 | |
19 | 2.5 M Sodium acetate | 0.1 M TRIS.HCl pH 8.5 | |
20 | 1.25 M Sodium acetate | 0.1 M TRIS.HCl pH 8.5 | |
21 | 1.2 M tri-Sodium citrate | 0.1 M TRIS.HCl pH 8.5 | |
22 | 0.6 M tri-Sodium citrate | 0.1 M TRIS.HCl pH 8.5 | |
23 | 0.6 M Sodium fluoride | 0.1 M TRIS.HCl pH 8.5 | |
24 | 0.3 M Sodium fluoride | 0.1 M TRIS.HCl pH 8.5 | |
25 | 3.5 M Sodium formate | 0.1 M Sodium acetate pH 4.6 | |
26 | 1.75 M Sodium formate | 0.1 M Sodium acetate pH 4.6 | |
27 | 3.5 M Sodium bromide | 0.1 M Sodium acetate pH 4.6 | |
28 | 1.75 M Sodium bromide | 0.1 M Sodium acetate pH 4.6 | |
29 | 2.4 M Sodium malonate | 0.1 M Sodium acetate pH 4.6 | |
30 | 1.2 M Sodium malonate | 0.1 M Sodium acetate pH 4.6 | |
31 | 3.5 M Sodium formate | 0.1 M MES pH 6.5 | |
32 | 1.75 M Sodium formate | 0.1 M MES pH 6.5 | |
33 | 3.5 M Sodium bromide | 0.1 M MES pH 6.5 | |
34 | 1.75 M Sodium bromide | 0.1 M MES pH 6.5 | |
35 | 2.4 M Sodium malonate | 0.1 M MES pH 6.5 | |
36 | 1.2 M Sodium malonate | 0.1 M MES pH 6.5 | |
37 | 3.5 M Sodium formate | 0.1 M HEPES pH 7.5 | |
38 | 1.75 M Sodium formate | 0.1 M HEPES pH 7.5 | |
39 | 3.5 M Sodium bromide | 0.1 M HEPES pH 7.5 | |
40 | 1.75 M Sodium bromide | 0.1 M HEPES pH 7.5 | |
41 | 2.4 M Sodium malonate | 0.1 M HEPES pH 7.5 | |
42 | 1.2 M Sodium malonate | 0.1 M HEPES pH 7.5 | |
43 | 3.5 M Sodium formate | 0.1 M TRIS.HCl pH 8.5 | |
44 | 1.75 M Sodium formate | 0.1 M TRIS.HCl pH 8.5 | |
45 | 3.5 M Sodium bromide | 0.1 M TRIS.HCl pH 8.5 | |
46 | 1.75 M Sodium bromide |
| 0.1 M TRIS.HCl pH 8.5 |
47 | 2.4 M Sodium malonate |
| 0.1 M TRIS.HCl pH 8.5 |
48 | 1.2 M Sodium malonate |
| 0.1 M TRIS.HCl pH 8.5 |
The Anions Suite
No | Precipitant | Salt | Buffer |
49 | 3.5 M Sodium nitrate | 0.1 M Sodium acetate pH 4.6 | |
50 | 1.75 M Sodium nitrate | 0.1 M Sodium acetate pH 4.6 | |
51 | 0.9 M Sodium phosphate | 0.9 M Potassium phosphate | 0.1 M Sodium acetate pH 4.6 |
52 | 0.45 M Sodium phosphate | 0.45 M Potassium phosphate | 0.1 M Sodium acetate pH 4.6 |
53 | 1.0 M Sodium succinate | 0.1 M Sodium acetate pH 4.6 | |
54 | 0.5 M Sodium succinate | 0.1 M Sodium acetate pH 4.6 | |
55 | 3.5 M Sodium nitrate | 0.1 M MES pH 6.5 | |
56 | 1.75 M Sodium nitrate | 0.1 M MES pH 6.5 | |
57 | 0.9 M Sodium phosphate | 0.9 M Potassium phosphate | 0.1 M MES pH 6.5 |
58 | 0.45 M Sodium phosphate | 0.45 M Potassium phosphate | 0.1 M MES pH 6.5 |
59 | 1.0 M Sodium succinate | 0.1 M MES pH 6.5 | |
60 | 0.5 M Sodium succinate | 0.1 M MES pH 6.5 | |
61 | 3.5 M Sodium nitrate | 0.1 M HEPES pH 7.5 | |
62 | 1.75 M Sodium nitrate | 0.1 M HEPES pH 7.5 | |
63 | 0.9 M Sodium phosphate | 0.9 M Potassium phosphate | 0.1 M HEPES pH 7.5 |
64 | 0.45 M Sodium phosphate | 0.45 M Potassium phosphate | 0.1 M HEPES pH 7.5 |
65 | 1.0 M Sodium succinate | 0.1 M HEPES pH 7.5 | |
66 | 0.5 M Sodium succinate | 0.1 M HEPES pH 7.5 | |
67 | 3.5 M Sodium nitrate | 0.1 M TRIS.HCl pH 8.5 | |
68 | 1.75 M Sodium nitrate | 0.1 M TRIS.HCl pH 8.5 | |
69 | 0.9 M Sodium phosphate | 0.9 M Potassium phosphate | 0.1 M TRIS.HCl pH 8.5 |
70 | 0.45 M Sodium phosphate | 0.45 M Potassium phosphate | 0.1 M TRIS.HCl pH 8.5 |
71 | 1.0 M Sodium succinate | 0.1 M TRIS.HCl pH 8.5 | |
72 | 0.5 M Sodium succinate | 0.1 M TRIS.HCl pH 8.5 | |
73 | 0.75 M Sodium sulfate | 0.1 M Sodium acetate pH 4.6 | |
74 | 0.37 M Sodium sulfate | 0.1 M Sodium acetate pH 4.6 | |
75 | 1.2 M K/Na tartrate | 0.1 M Sodium tartrate pH 5.6 | |
76 | 0.6 M K/Na tartrate | 0.1 M Sodium tartrate pH 5.6 | |
77 | 2.4 M Sodium thiocyanate | 0.1 M Sodium acetate pH 4.6 | |
78 | 1.2 M Sodium thiocyanate | 0.1 M Sodium acetate pH 4.6 | |
79 | 0.75 M Sodium sulfate | 0.1 M MES pH 6.5 | |
80 | 0.37 M Sodium sulfate | 0.1 M MES pH 6.5 | |
81 | 1.2 M K/Na tartrate | 0.1 M MES pH 6.5 | |
82 | 0.6 M K/Na tartrate | 0.1 M MES pH 6.5 | |
83 | 2.4 M Sodium thiocyanate | 0.1 M MES pH 6.5 | |
84 | 1.2 M Sodium thiocyanate | 0.1 M MES pH 6.5 | |
85 | 0.75 M Sodium sulfate | 0.1 M HEPES pH 7.5 | |
86 | 0.37 M Sodium sulfate | 0.1 M HEPES pH 7.5 | |
87 | 1.2 M K/Na tartrate | 0.1 M HEPES pH 7.5 | |
88 | 0.6 M K/Na tartrate | 0.1 M HEPES pH 7.5 | |
89 | 2.4 M Sodium thiocyanate | 0.1 M HEPES pH 7.5 | |
90 | 1.2 M Sodium thiocyanate | 0.1 M HEPES pH 7.5 | |
91 | 0.75 M Sodium sulfate | 0.1 M TRIS.HCl pH 8.5 | |
92 | 0.37 M Sodium sulfate | 0.1 M TRIS.HCl pH 8.5 | |
93 | 1.2 M K/Na tartrate | 0.1 M TRIS.HCl pH 8.5 | |
94 | 0.6 M K/Na tartrate | 0.1 M TRIS.HCl pH 8.5 | |
95 | 2.4 M Sodium thiocyanate | 0.1 M TRIS.HCl pH 8.5 | |
96 | 1.2 M Sodium thiocyanate | 0.1 M TRIS.HCl pH 8.5 |
The Nucleix Suite
No | Precipitant | Salt 2 | Salt 3 | Buffer |
1 | 2 M Lithium sulfate |
| 0.05 M MES pH 5.6 | |
2 | 2.5 M Ammonium sulfate |
| 0.05 M MES pH 5.6 | |
3 | 20%(v/v) MPD |
| 0.05 M MES pH 5.6 | |
4 | 10%(v/v) PEG 400 | 0.01 M Magnesium sulfate | 0.05 M MES pH 5.6 | |
5 | 5%(w/v) PEG 8000 | 0.01 M Magnesium chloride | 0.05 M MES pH 5.6 | |
6 | 20%(w/v) PEG 8000 | 0.01 M Magnesium chloride | 0.05 M MES pH 5.6 | |
7 | 15%(v/v) Isopropanol |
| 0.05 M MES pH 6.0 | |
8 | 0.6 M Sodium chloride | 0.005 M Magnesium sulfate | 0.05 M MES pH 6.0 | |
9 | 10%(v/v) PEG 400 | 0.01 M Magnesium chloride | 0.05 M MES pH 6.0 | |
10 | 5%(w/v) PEG 4000 |
| 0.05 M MES pH 6.0 | |
11 | 1 M Lithium sulfate |
| 0.05 M Sodium cacodylate pH 6.0 | |
12 | 1.8 M Lithium sulfate |
| 0.05 M Sodium cacodylate pH 6.0 | |
13 | 1.7 M Ammonium sulfate |
| 0.05 M Sodium cacodylate pH 6.0 | |
14 | 15%(v/v) Isopropanol | 0.025 M Magnesium chloride | 0.05 M Sodium cacodylate pH 6.0 | |
15 | 5%(v/v) MPD |
| 0.05 M Sodium cacodylate pH 6.0 | |
16 | 30%(v/v) MPD |
| 0.05 M Sodium cacodylate pH 6.0 | |
17 | 10%(w/v) PEG 4000 | 0.01 M Calcium chloride | 0.05 M Sodium cacodylate pH 6.0 | |
18 | 1.3 M Lithium sulfate |
| 0.05 M Sodium cacodylate pH 6.5 | |
19 | 2 M Ammonium sulfate |
| 0.05 M Sodium cacodylate pH 6.5 | |
20 | 10%(v/v) Isopropanol | 0.015 M Magnesium acetate | 0.05 M Sodium cacodylate pH 6.5 | |
21 | 10%(w/v) 1,6-Hexanediol | 0.005 M Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
22 | 15%(v/v) PEG 400 |
| 0.05 M Sodium cacodylate pH 6.5 | |
23 | 10%(w/v) PEG 4000 | 0.01 M Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
24 | 10%(w/v) PEG 4000 | 0.01 M Calcium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
25 | 30%(w/v) PEG 4000 |
| 0.05 M Sodium cacodylate pH 6.5 | |
26 | 10%(w/v) PEG 8000 | 0.1 M Magnesium acetate | 0.05 M Sodium cacodylate pH 6.5 | |
27 | 30%(w/v) PEG 8000 | 0.01 M Magnesium acetate | 0.05 M Sodium cacodylate pH 6.5 | |
28 | 1.6 M Lithium sulfate |
| 0.05 M HEPES Sodium salt pH 7.0 | |
29 | 4 M Lithium chloride |
| 0.05 M HEPES Sodium salt pH 7.0 | |
30 | 1.6 M Ammonium sulfate |
| 0.05 M HEPES Sodium salt pH 7.0 | |
31 | 25%(v/v) PEG MME 550 |
| 0.05 M HEPES Sodium salt pH 7.0 | |
32 | 20%(w/v) 1,6-Hexanediol | 0.01 M Magnesium chloride | 0.05 M HEPES Sodium salt pH 7.0 | |
33 | 30%(w/v) 1,6-Hexanediol | 0.01 M Magnesium chloride | 0.05 M HEPES Sodium salt pH 7.0 | |
34 | 15%(v/v) MPD | 0.005 M Magnesium sulfate | 0.05 M HEPES Sodium salt pH 7.0 | |
35 | 5%(v/v) PEG 400 | 0.01 M Magnesium chloride | 0.05 M HEPES Sodium salt pH 7.0 | |
36 | 10%(v/v) PEG 400 | 0.01 M Calcium chloride | 0.05 M HEPES Sodium salt pH 7.0 | |
37 | 20%(v/v) PEG 200 | 0.025 M Magnesium sulfate | 0.05 M HEPES Sodium salt pH 7.0 | |
38 | 5%(w/v) PEG 4000 | 0.15 M Magnesium acetate | 0.05 M HEPES Sodium salt pH 7.0 | |
39 | 5%(w/v) PEG 8000 | 0.02 M Magnesium chloride | 0.05 M HEPES Sodium salt pH 7.0 | |
40 | 1.6 M Ammonium sulfate |
| 0.05 M TRIS.HCl pH 7.5 | |
41 | 10%(v/v) PEG MME 550 | 0.015 M Magnesium chloride | 0.05 M TRIS.HCl pH 7.5 | |
42 | 5%(v/v) Isopropanol |
| 0.05 M TRIS.HCl pH 7.5 | |
43 | 10%(v/v) MPD | 0.01 M Magnesium chloride | 0.05 M TRIS.HCl pH 7.5 | |
44 | 10%(w/v) PEG 4000 | 0.05 M Magnesium chloride | 0.05 M TRIS.HCl pH 7.5 | |
45 | 1.8 M Ammonium sulfate |
| 0.05 M TRIS.HCl pH 8.5 | |
46 | 35%(w/v) 1,6-Hexanediol |
| 0.05 M TRIS.HCl pH 8.5 | |
47 | 30%(v/v) PEG 400 | 0.01 M Magnesium chloride | 0.05 M TRIS.HCl pH 8.5 | |
48 | 30%(w/v) PEG 4000 | 0.01 M Calcium chloride |
| 0.05 M TRIS.HCl pH 8.5 |
49 |
| 80 mM Magnesium chloride |
| 0.05 M HEPES pH 7.5 |
The Nucleix Suite
50 | 9%(v/v) Isopropanol | 18 mM Magnesium chloride | 1 mM Copper sulfate | 0.05 M Sodium cacodylate pH 6.0 |
51 | 9%(v/v) Isopropanol | 18 mM Magnesium chloride | 1.8 mM [Co(NH3)6]Cl3 | 0.05 M Sodium cacodylate pH 6.5 |
52 | 9%(v/v) Isopropanol | 18 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
53 | 4.5%(v/v) MPD | 18 mM Magnesium chloride | 0.9 mM [Co(NH3)6]Cl3 | 0.05 M Sodium cacodylate pH 7.0 |
54 | 5%(v/v) PEG 400 | 36 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
55 | 10%(v/v) Isopropanol | 10 mM Magnesium chloride | 0.05 M Sodium succinate pH 5.5 | |
56 | 15%(v/v) Ethanol | 20 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.0 | |
57 | 15%(v/v) Ethanol | 20 mM Magnesium chloride | 1.0 mM [Co(NH3)6]Cl3 | 0.05 M Sodium cacodylate pH 7.0 |
58 | 10%(v/v) tert-Butanol | 5 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 7.0 | |
59 | 5%(v/v) PEG 400 | 30 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 7.0 | |
60 | 5%(v/v) Isopropanol | 100 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
61 | 20%(v/v) Ethanol | 10 mM Magnesium chloride | 0.05 M TRIS pH 8.0 | |
62 | 5%(w/v) PEG 8000 | 20 mM Magnesium chloride | 0.05 M HEPES pH 7.5 | |
63 | 5%(w/v) PEG 4000 | 20 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.0 | |
64 | 10%(v/v) Isopropanol | 10 mM Magnesium chloride | 5 mM Calcium chloride | 0.05 M Sodium cacodylate pH 6.0 |
65 | 5%(v/v) PEG 400 | 9 mM Magnesium chloride | 1.8 mM [Co(NH3)6]Cl3 | 0.05 M Sodium cacodylate pH 7.0 |
0.9 mM Spermidine | ||||
66 | 10%(v/v) Isopropanol | 10 mM Magnesium chloride | 1 mM Copper sulfate | 0.05 M Sodium cacodylate pH 6.5 |
67 | 10%(v/v) 1,5-Hexandiol | 20 mM Magnesium chloride | 2 mM Calcium chloride | 0.05 M Sodium cacodylate pH 6.0 |
68 | 10%(v/v) Dioxane | 15 mM Magnesium chloride | 0.05 M HEPES pH 7.5 | |
69 | 10%(v/v) PEG 400 | 15 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.0 | |
70 | 9%(v/v) Isopropanol | 2.5 mM Spermine | 0.05 M Sodium cacodylate pH 6.5 | |
71 | 2.0 mM Spermine | 80 mM Calcium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
72 | 5 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | ||
73 | 1.3 M Lithium sulfate | 30 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
74 | 5%(v/v) Isopropanol | 0.05 M Sodium cacodylate pH 6.0 | ||
75 | 10%(v/v) Ethanol | 100 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
76 | 2.5 M Sodium chloride | 10 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.0 | |
77 | 5%(v/v) Isopropanol | 10 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
78 | 2.0 M Lithium sulfate | 15 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
79 | 2.0 M Ammonium sulfate | 20 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
80 | 3.0 M Ammonium sulfate | 10 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
81 | 1.0 M Ammonium sulfate | 15 mM Magnesium chloride | 0.05 M HEPES pH 7.5 | |
82 | 2.5 M Sodium chloride | 0.05 M Sodium cacodylate pH 6.0 | ||
83 | 2.0 M Lithium chloride | 200 mM Calcium acetate | 0.05 M Sodium cacodylate pH 6.0 | |
84 | 2.0 M Sodium chloride | 15 mM Magnesium chloride | 1.0 mM [Co(NH3)6]Cl3 | 0.05 M Sodium cacodylate pH 6.5 |
85 | 20%(w/v) PEG 1000 | 200 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
86 | 1.0 M Sodium tartrate |
| 0.05 M TRIS pH 7.5 | |
87 | 2.5 M Sodium chloride |
| 0.05 M TRIS pH 7.5 | |
88 | 2.5 M Potassium chloride |
| 0.05 M Sodium cacodylate pH 6.0 | |
89 | 15%(v/v) Ethanol |
| 0.05 M TRIS pH 8.0 | |
90 | 2.0 M Lithium sulfate | 15 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.0 | |
91 | 25%(v/v) MPD | 20 mM Magnesium acetate | 100 mM Sodium chloride | 0.05 M Sodium cacodylate pH 6.0 |
92 | 3.0 M Ammonium sulfate | 20 mM Magnesium chloride | 0.05 M Sodium succinate pH 5.5 | |
93 | 2.5 M Potassium chloride | 0.05 M Sodium cacodylate pH 6.5 | ||
94 | 1.5 M Lithium sulfate | 50 mM Magnesium chloride | 0.05 M Sodium cacodylate pH 6.5 | |
95 | 2.0 M Lithium chloride | 1.0 mM Spermine | 30 mM Calcium chloride | 0.05 M Sodium cacodylate pH 6.5 |
96 | 10 mM Magnesium chloride |
| 0.05 M Sodium cacodylate pH 6.5 |
The PACT Suit
Number | Precipitant | Salt | Buffer |
| ||
1 | 25% w/v PEG 1500 | 0.1 M SPG buffer pH 4 |
| |||
2 | 25% w/v PEG 1500 | 0.1 M SPG buffer pH 5 |
| |||
3 | 25% w/v PEG 1500 | 0.1 M SPG buffer pH 6 |
| |||
4 | 25% w/v PEG 1500 | 0.1 M SPG buffer pH 7 |
| |||
5 | 25% w/v PEG 1500 | 0.1 M SPG buffer pH 8 |
| |||
6 | 25% w/v PEG 1500 | 0.1 M SPG buffer pH 9 |
| |||
7 | 20% w/v PEG 6000 | 0.2 M Sodium chloride | 0.1 M Sodium acetate pH 5 |
| ||
8 | 20% w/v PEG 6000 | 0.2 M Ammonium chloride | 0.1 M Sodium acetate pH 5 |
| ||
9 | 20% w/v PEG 6000 | 0.2 M Lithium chloride | 0.1 M Sodium acetate pH 5 |
| ||
10 | 20% w/v PEG 6000 | 0.2 M Magnesium chloride | 0.1 M Sodium acetate pH 5 |
| ||
11 | 20% w/v PEG 6000 | 0.2 M Calcium chloride | 0.1 M Sodium acetate pH 5 |
| ||
12 | 20% w/v PEG 6000 | 0.01 M Zinc chloride | 0.1 M Sodium acetate pH 5 |
| ||
13 | 25% w/v PEG 1500 | 0.1 M MIB buffer pH 4 |
| |||
14 | 25% w/v PEG 1500 | 0.1 M MIB buffer pH 5 |
| |||
15 | 25% w/v PEG 1500 | 0.1 M MIB buffer pH 6 |
| |||
16 | 25% w/v PEG 1500 | 0.1 M MIB buffer pH 7 |
| |||
17 | 25% w/v PEG 1500 | 0.1 M MIB buffer pH 8 |
| |||
18 | 25% w/v PEG 1500 | 0.1 M MIB buffer pH 9 |
| |||
19 | 20% w/v PEG 6000 | 0.2 M Sodium chloride | 0.1 M MES pH 6 |
| ||
20 | 20% w/v PEG 6000 | 0.2 M Ammonium chloride | 0.1 M MES pH 6 |
| ||
21 | 20% w/v PEG 6000 | 0.2 M Lithium chloride | 0.1 M MES pH 6 |
| ||
22 | 20% w/v PEG 6000 | 0.2 M Magnesium chloride | 0.1 M MES pH 6 |
| ||
23 | 20% w/v PEG 6000 | 0.2 M Calcium chloride | 0.1 M MES pH 6 |
| ||
24 | 20% w/v PEG 6000 | 0.01 M Zinc chloride | 0.1 M MES pH 6 |
| ||
25 | 25% w/v PEG 1500 | 0.1 M PCB buffer pH 4 |
| |||
26 | 25% w/v PEG 1500 | 0.1 M PCB buffer pH 5 |
| |||
27 | 25% w/v PEG 1500 | 0.1 M PCB buffer pH 6 |
| |||
28 | 25% w/v PEG 1500 | 0.1 M PCB buffer pH 7 |
| |||
29 | 25% w/v PEG 1500 | 0.1 M PCB buffer pH 8 |
| |||
30 | 25% w/v PEG 1500 | 0.1 M PCB buffer pH 9 |
| |||
31 | 20% w/v PEG 6000 | 0.2 M Sodium chloride | 0.1 M HEPES pH 7 |
| ||
32 | 20% w/v PEG 6000 | 0.2 M Ammonium chloride | 0.1 M HEPES pH 7 |
| ||
33 | 20% w/v PEG 6000 | 0.2 M Lithium chloride | 0.1 M HEPES pH 7 |
| ||
34 | 20% w/v PEG 6000 | 0.2 M Magnesium chloride | 0.1 M HEPES pH 7 |
| ||
35 | 20% w/v PEG 6000 | 0.2 M Calcium chloride | 0.1 M HEPES pH 7 |
| ||
36 | 20% w/v PEG 6000 | 0.01 M Zinc chloride | 0.1 M HEPES pH 7 |
| ||
37 | 25% w/v PEG 1500 | 0.1 M MMT buffer pH 4 |
| |||
38 | 25% w/v PEG 1500 | 0.1 M MMT buffer pH 5 |
| |||
39 | 25% w/v PEG 1500 | 0.1 M MMT buffer pH 6 |
| |||
40 | 25% w/v PEG 1500 | 0.1 M MMT buffer pH 7 |
| |||
41 | 25% w/v PEG 1500 | 0.1 M MMT buffer pH 8 |
| |||
42 | 25% w/v PEG 1500 | 0.1 M MMT buffer pH 9 |
| |||
43 | 20% w/v PEG 6000 | 0.2 M Sodium chloride | 0.1 M Tris pH 8 |
| ||
44 | 20% w/v PEG 6000 | 0.2 M Ammonium chloride | 0.1 M Tris pH 8 |
| ||
45 | 20% w/v PEG 6000 | 0.2 M Lithium chloride | 0.1 M Tris pH 8 |
| ||
46 | 20% w/v PEG 6000 | 0.2 M Magnesium chloride | 0.1 M Tris pH 8 |
| ||
47 | 20% w/v PEG 6000 | 0.2 M Calcium chloride | 0.1 M Tris pH 8 |
| ||
48 | 20% w/v PEG 6000 | 0.1 M Tris pH 8 |
| |||
49 | 20% w/v PEG 3350 | 0.2 M Sodium fluoride |
|
| ||
50 | 20% w/v PEG 3350 | 0.2 M Sodium bromide | ||||
51 | 20% w/v PEG 3350 | 0.2 M Sodium iodide | ||||
52 | 20% w/v PEG 3350 | 0.2 M Potassium thiocyanate | ||||
53 | 20% w/v PEG 3350 | 0.2 M Sodium nitrate | ||||
54 | 20% w/v PEG 3350 | 0.2 M Sodium formate | ||||
55 | 20% w/v PEG 3350 | 0.2 M Sodium acetate | ||||
56 | 20% w/v PEG 3350 | 0.2 M Sodium sulphate | ||||
57 | 20% w/v PEG 3350 | 0.2 M Potassium/sodium tartrate | ||||
58 | 20% w/v PEG 3350 | 0.2 M Sodium/potassium phosphate | ||||
59 | 20% w/v PEG 3350 | 0.2 M Sodium citrate | ||||
60 | 20% w/v PEG 3350 | 0.2 M Sodium malonate | ||||
61 | 20% w/v PEG 3350 | 0.2 M Sodium fluoride | 0.1 M Bis Tris propane pH 6.5 | |||
62 | 20% w/v PEG 3350 | 0.2 M Sodium bromide | 0.1 M Bis Tris propane pH 6.5 | |||
63 | 20% w/v PEG 3350 | 0.2 M Sodium iodide | 0.1 M Bis Tris propane pH 6.5 | |||
64 | 20% w/v PEG 3350 | 0.2 M Potassium thiocyanate | 0.1 M Bis Tris propane pH 6.5 | |||
65 | 20% w/v PEG 3350 | 0.2 M Sodium nitrate | 0.1 M Bis Tris propane pH 6.5 | |||
66 | 20% w/v PEG 3350 | 0.2 M Sodium formate | 0.1 M Bis Tris propane pH 6.5 | |||
67 | 20% w/v PEG 3350 | 0.2 M Sodium acetate | 0.1 M Bis Tris propane pH 6.5 | |||
68 | 20% w/v PEG 3350 | 0.2 M Sodium sulphate | 0.1 M Bis Tris propane pH 6.5 | |||
69 | 20% w/v PEG 3350 | 0.2 M Potassium/sodium tartrate | 0.1 M Bis Tris propane pH 6.5 | |||
70 | 20% w/v PEG 3350 | 0.2 M Sodium/potassium phosphate | 0.1 M Bis Tris propane pH 6.5 | |||
71 | 20% w/v PEG 3350 | 0.2 M Sodium citrate | 0.1 M Bis Tris propane pH 6.5 | |||
72 | 20% w/v PEG 3350 | 0.2 M Sodium malonate | 0.1 M Bis Tris propane pH 6.5 | |||
73 | 20% w/v PEG 3350 | 0.2 M Sodium fluoride | 0.1 M Bis Tris propane pH 7.5 | |||
74 | 20% w/v PEG 3350 | 0.2 M Sodium bromide | 0.1 M Bis Tris propane pH 7.5 | |||
75 | 20% w/v PEG 3350 | 0.2 M Sodium iodide | 0.1 M Bis Tris propane pH 7.5 | |||
76 | 20% w/v PEG 3350 | 0.2 M Potassium thiocyanate | 0.1 M Bis Tris propane pH 7.5 | |||
77 | 20% w/v PEG 3350 | 0.2 M Sodium nitrate | 0.1 M Bis Tris propane pH 7.5 | |||
78 | 20% w/v PEG 3350 | 0.2 M Sodium formate | 0.1 M Bis Tris propane pH 7.5 | |||
79 | 20% w/v PEG 3350 | 0.2 M Sodium acetate | 0.1 M Bis Tris propane pH 7.5 | |||
80 | 20% w/v PEG 3350 | 0.2 M Sodium sulphate | 0.1 M Bis Tris propane pH 7.5 | |||
81 | 20% w/v PEG 3350 | 0.2 M Potassium/sodium tartarte | 0.1 M Bis Tris propane pH 7.5 | |||
82 | 20% w/v PEG 3350 | 0.2 M Sodium/potassium phosphate | 0.1 M Bis Tris propane pH 7.5 | |||
83 | 20% w/v PEG 3350 | 0.2 M Sodium citrate | 0.1 M Bis Tris propane pH 7.5 | |||
84 | 20% w/v PEG 3350 | 0.2 M Sodium malonate | 0.1 M Bis Tris propane pH 7.5 | |||
85 | 20% w/v PEG 3350 | 0.2 M Sodium fluoride | 0.1 M Bis Tris propane pH 8.5 | |||
86 | 20% w/v PEG 3350 | 0.2 M Sodium bromide | 0.1 M Bis Tris propane pH 8.5 | |||
87 | 20% w/v PEG 3350 | 0.2 M Sodium iodide | 0.1 M Bis Tris propane pH 8.5 | |||
88 | 20% w/v PEG 3350 | 0.2 M Potassium thiocyanate | 0.1 M Bis Tris propane pH 8.5 | |||
89 | 20% w/v PEG 3350 | 0.2 M Sodium nitrate | 0.1 M Bis Tris propane pH 8.5 | |||
90 | 20% w/v PEG 3350 | 0.2 M Sodium formate | 0.1 M Bis Tris propane pH 8.5 | |||
91 | 20% w/v PEG 3350 | 0.2 M Sodium acetate | 0.1 M Bis Tris propane pH 8.5 | |||
92 | 20% w/v PEG 3350 | 0.2 M Sodium sulphate | 0.1 M Bis Tris propane pH 8.5 | |||
93 | 20% w/v PEG 3350 | 0.2 M Potassium/sodium tartrate | 0.1 M Bis Tris propane pH 8.5 | |||
94 | 20% w/v PEG 3350 | 0.2 M Sodium/potassium phosphate | 0.1 M Bis Tris propane pH 8.5 | |||
95 | 20% w/v PEG 3350 | 0.2 M Sodium citrate | 0.1 M Bis Tris propane pH 8.5 | |||
96 | 20% w/v PEG 3350 | 0.2 M Sodium malonate | 0.1 M Bis Tris propane pH 8.5 | |||
The Protein Complex Suite
No. | Precipitant 1 | Precipitant 2 | Salt | Buffer |
1 | 25%(w/v) PEG 350 MME | 0.1 M TRIS pH 8 | ||
2 | 15%(v/v) PEG 400 | 0.1 Calcium acetate | 0.1 M MES pH 6 | |
3 | 20%(v/v) PEG 400 | 0.1 Lithium chloride | 0.1 M HEPES pH 7.5 | |
4 | 25%(v/v) PEG 400 | 0.1 M TRIS pH 8 | ||
5 | 15%(w/v) PEG 550 MME | 0.1 M MES pH 6.5 | ||
6 | 25%(w/v) PEG 1000 | 0.2 M Sodium chloride | 0.1 M Na/K phosphate pH 6.5 | |
7 | 20%(w/v) PEG 1500 | 0.1 M Ammonium sulfate | 0.1 M TRIS pH 7.5 | |
8 | 10%(w/v) PEG 2000 MME | 0.2 M Ammonium sulfate | 0.1 M Sodium acetate pH 5.5 | |
9 | 20%(w/v) PEG 2000 MME | 0.2 M Sodium chloride | 0.1 M MES pH 6 | |
10 | 15%(w/v) PEG 2000 MME | 0.1 M Potassium chloride | 0.1 M TRIS pH 8 | |
11 | 25%(w/v) PEG 2000 MME | 0.1 M HEPES pH 7.5 | ||
12 | 5%(w/v) PEG 4000 | 0.2 M Sodium acetate | 0.1 M Sodium citrate pH 5.5 | |
13 | 5%(w/v) PEG 4000 | 0.2 M Lithium sulfate | 0.1 M TRIS pH 7.5 | |
14 | 10%(w/v) PEG 4000 | 0.1 M Calcium acetate | 0.1 M Sodium acetate pH 4.5 | |
15 | 10%(w/v) PEG 4000 | 0.2 M Sodium acetate | 0.1 M Sodium citrate pH 5.5 | |
16 | 10%(w/v) PEG 4000 | 0.2 M Sodium chloride | 0.1 M MES pH 6.5 | |
17 | 10%(w/v) PEG 4000 | 0.1 M Magnesium chloride | 0.1 M HEPES pH 7.5 | |
18 | 10%(w/v) PEG 4000 | 10% (v/v) Isopropanol | 0.1 M HEPES pH 7 | |
19 | 15%(w/v) PEG 4000 | 0.2 M Ammonium acetate | 0.1 M Sodium acetate pH 4 | |
20 | 15%(w/v) PEG 4000 | 0.1 M Magnesium chloride | 0.1 M Sodium citrate pH 5 | |
21 | 15%(w/v) PEG 4000 | 0.1 M Sodium cacodylate pH 6 | ||
22 | 15%(w/v) PEG 4000 | 0.15 M Ammonium sulfate | 0.1 M MES pH 6 | |
23 | 15%(w/v) PEG 4000 | 0.1 M HEPES pH 7 | ||
24 | 15%(w/v) PEG 4000 | 0.1 M Magnesium chloride | 0.1 M HEPES pH 7 | |
25 | 15%(w/v) PEG 4000 | 0.15 M Ammonium sulfate | 0.1 M TRIS pH 8 | |
26 | 20%(w/v) PEG 4000 | 0.1 M Sodium citrate pH 4.5 | ||
27 | 20%(w/v) PEG 4000 | 0.2 M Ammonium acetate | 0.1 M Sodium acetate pH 5 | |
28 | 20%(w/v) PEG 4000 | 0.2 M Lithium sulfate | 0.1 M MES pH 6 | |
29 | 20%(w/v) PEG 4000 | 0.1 M TRIS pH 8 | ||
30 | 20%(w/v) PEG 4000 | 0.15 M Ammonium sulfate | 0.1 M HEPES pH 7 | |
31 | 20%(w/v) PEG 4000 | 20% (v/v) Isopropanol | 0.1 M Sodium citrate pH 5.6 | |
32 | 20%(w/v) PEG 4000 | 0.2 M Sodium chloride | 0.1 M TRIS pH 8 | |
33 | 25%(w/v) PEG 4000 | 0.1 M Sodium cacodylate pH 5.5 | ||
34 | 25%(w/v) PEG 4000 | 0.15 M Ammonium sulfate | 0.1 M MES pH 5.5 | |
35 | 25%(w/v) PEG 4000 | 0.1 M Sodium cacodylate pH 6.5 | ||
36 | 25%(w/v) PEG 4000 | 0.2 M Potassium iodide | 0.1 M MES pH 6.5 | |
37 | 25%(w/v) PEG 4000 | 0.2 M Sodium chloride | 0.1 M HEPES pH 7.5 | |
38 | 10%(w/v) PEG 5000 MME | 12% (v/v) 1-propanol | 0.1 M MES pH 6.5 | |
39 | 15%(w/v) PEG 5000 MME | 0.1 M Potassium chloride | 0.1 M HEPES pH 7 | |
40 | 20%(w/v) PEG 5000 MME | 0.2 M Ammonium sulfate | 0.1 M TRIS pH 7.5 | |
41 | 8%(w/v) PEG 6000 | 0.1 M Magnesium chloride | 0.1 M MES pH 6 | |
42 | 8%(w/v) PEG 6000 | 0.15 M Sodium chloride | 0.1 M TRIS pH 8 | |
43 | 15%(w/v) PEG 6000 | 0.1 M Sodium citrate pH 5.5 | ||
44 | 15%(w/v) PEG 6000 | 0.1 M Magnesium acetate | 0.1 M Sodium cacodylate pH 6.5 | |
45 | 15%(w/v) PEG 6000 | 5%(v/v) MPD | 0.1 M MES pH 6.5 | |
46 | 15%(w/v) PEG 6000 | 0.1 M Potassium chloride | 0.1 M HEPES pH 7.5 | |
47 | 15%(w/v) PEG 6000 | 0.1 M TRIS pH 7.5 | ||
48 | 20%(w/v) PEG 6000 | 0.1 M TRIS pH 8.5 | ||
The Protein Complex Suite
Number | Precipitant 1 | Precipitant 2 | Salt | Buffer |
49 | 8%(w/v) PEG 8000 | 0.1 M Magnesium acetate | 0.1 M Sodium acetate pH 4.5 | |
50 | 8%(w/v) PEG 8000 | 0.1 M Sodium citrate pH 5 | ||
51 | 8%(w/v) PEG 8000 | 0.2 M Sodium chloride | 0.1 M Sodium cacodylate pH 6 | |
52 | 8%(w/v) PEG 8000 | 0.1 M HEPES pH 7 | ||
53 | 8%(w/v) PEG 8000 | 0.1 M TRIS pH 8 | ||
54 | 12%(w/v) PEG 8000 | 0.1 M Calcium acetate | 0.1 M Sodium cacodylate pH 5.5 | |
55 | 12%(w/v) PEG 8000 |
|
| 0.1 M NaH2PO4 p |
56 | 12%(w/v) PEG 8000 | 0.1 M Magnesium acetate | 0.1 M MOPS pH 7.5 | |
57 | 12%(w/v) PEG 8000 | 0.2 M Sodium chloride | 0.1 M HEPES pH 7.5 | |
58 | 12%(w/v) PEG 8000 | 0.2 M Ammonium sulfate | 0.1 M TRIS pH 8.5 | |
59 | 20%(w/v) PEG 8000 | 0.1 M Sodium citrate pH 5 | ||
60 | 20%(w/v) PEG 8000 | 0.2 M Ammonium sulfate | 0.1 M MES pH 6.5 | |
61 | 20%(w/v) PEG 8000 | 0.1 M HEPES pH 7 | ||
62 | 20%(w/v) PEG 8000 | 0.2 M Lithium chloride | 0.1 M TRIS pH 8 | |
63 | 10%(w/v) PEG 10000 | 0.1 M Magnesium acetate | 0.1 M MES pH 6.5 | |
64 | 18%(w/v) PEG 12000 | 0.1 M HEPES pH 7 | ||
65 | 8%(w/v) PEG 20000 | 0.1 M Sodium chloride | 0.1 M TRIS pH 8 | |
66 | 15%(w/v) PEG 20000 | 0.1 M HEPES pH 7 | ||
67 | 0.5 M Ammonium sulfate | 0.1 M MES pH 6.5 | ||
68 | 1 M Ammonium sulfate | 0.1 M Sodium acetate pH 5 | ||
69 | 1 M Ammonium sulfate | 0.1 M MES pH 6.5 | ||
70 | 1 M Ammonium sulfate | 0.1 M TRIS pH 8 | ||
71 | 1.5 M Ammonium sulfate | 0.1 M Sodium acetate pH 5 | ||
72 | 1.5 M Ammonium sulfate | 0.1 M HEPES pH 7 | ||
73 | 1.5 M Ammonium sulfate | 0.1 M TRIS pH 8 | ||
74 | 2 M Ammonium sulfate | 0.1 M Sodium acetate pH 5 | ||
75 | 2 M Ammonium sulfate | 0.1 M HEPES pH 7 | ||
76 | 2 M Ammonium sulfate | 0.1 M TRIS pH 8 | ||
77 | 1 M Ammonium sulfate | 1 M Potassium chloride | 0.1 M HEPES pH 7 | |
78 | 2 M Sodium formate | 0.1 M Sodium acetate pH 5 | ||
79 | 3 M Sodium formate | 0.1 M TRIS pH 7.5 | ||
80 | 0.8 M Potassium |
|
| Sodium phosphate pH 7.5 |
81 | 1.3 M Potassium |
|
| Sodium phosphate pH 7.0 |
82 | 1.6 M Potassium |
|
| Sodium phosphate pH 6.5 |
83 | 1 M Sodium acetate | 0.1 M HEPES pH 7.5 | ||
84 | 1 M Sodium citrate | 0.1 M HEPES pH 7 | ||
85 | 2 M Sodium chloride | 0.1 M Sodium citrate pH 6 | ||
86 | 1 M Lithium sulfate | 0.1 M MES pH 6.5 | ||
87 | 1.6 M Lithium sulfate | 0.1 M TRIS pH 8 | ||
88 | 1.4 M Sodium malonate | pH 6.0 | ||
89 | 1.2 M Na/K tartrate |
|
| 0.1 M TRIS pH 8 |
90 | 1.6 M Magnesium sulfate | 0.1 M MES pH 6.5 | ||
91 | 15%(v/v) MPD | 2%(w/v) PEG 4000 | 0.1 M Sodium acetate pH 5 | |
92 | 25%(v/v) MPD | 0.05 M Calcium acetate | 0.1 M Sodium cacodylate pH 6 | |
93 | 50%(v/v) MPD | 0.1 M Imidazole pH 7 | ||
94 | 10%(v/v) Isopropanol | 5%(w/v) PEG 4000 | 0.05 M MgCl2 | 0.1 M MES pH 6.5 |
95 | 25%(v/v) Isopropanol | 0.2 M Ammonium acetate | 0.1 M HEPES pH 7.5 | |
96 | 15%(v/v) Ethanol | 5%(v/v) MPD | 0.1 M Sodium chloride | 0.1 M TRIS pH 8 |
Natrix HT (HR2-131)
No | Salt | Buffer | Precipitant |
1 | 0.01 M Magnesium chloride hexahydrate | 0.05 M MES monohydrate pH 5.6 | 1.8 M Lithium sulfate monohydrate |
2 | 0.01 M Magnesium acetate tetrahydrate | 0.05 M MES monohydrate pH 5.6 | 2.5 M (NH4)2SO4 |
3 | 0.1 M Magnesium acetate tetrahydrate | 0.05 M MES monohydrate pH 5.6 | 20% v/v MPD |
4 | 0.2 M KCl 0.01 M Magnesium sulfate heptahydrate | 0.05 M MES monohydrate pH 5.6 | 10% v/v PEG 400 |
5 | 0.2 M KCl 0.01 M Magnesium chloride hexahydrate | 0.05 M MES monohydrate pH 5.6 | 5% w/v PEG 8000 |
6 | 0.1 M (NH4)2SO4 0.01 M Magnesium chloride hexahydrate | 0.05 M MES monohydrate pH 5.6 | 20% w/v PEG 8000 |
7 | 0.02 M Magnesium chloride hexahydrate | 0.05 M MES monohydrate pH 6.0 | 15% v/v 2-Propanol |
8 | 0.1 M NH4CH3CO2 0.005 M Magnesium sulfate heptahydrate | 0.05 M MES monohydrate pH 6.0 | 0.6 M NaCl |
9 | 0.1 M KCl 0.01 M Magnesium chloride hexahydrate | 0.05 M MES monohydrate pH 6.0 | 10% v/v PEG 400 |
10 | 0.005 M Magnesium sulfate heptahydrate | 0.05 M MES monohydrate pH 6.0 | 5% w/v PEG 4000 |
11 | 0.01 M Magnesium chloride hexahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 1.0 M Lithium sulfate monohydrate |
12 | 0.01 M Magnesium sulfate heptahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 1.8 M Lithium sulfate monohydrate |
13 | 0.015 M Magnesium acetate tetrahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 1.7 M (NH4)2SO4 |
14 | 0.1 M KCl 0.025 M Magnesium chloride hexahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 15% v/v 2-Propanol |
15 | 0.04 M Magnesium chloride hexahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 5% v/v MPD |
16 | 0.04 M Magnesium acetate tetrahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 30% v/v MPD |
17 | 0.2 M KCl 0.01 M Calcium chloride dihydrate | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 10% w/v PEG 4000 |
18 | 0.01 M Magnesium acetate tetrahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.5 | 1.3 M Lithium sulfate monohydrate |
19 | 0.01 M Magnesium sulfate heptahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.5 | 2.0 M (NH4)2SO4 |
20 | 0.1 M NH4CH3CO2 0.015 M Mg(CH3COO)2 • 4H2O | 0.05 M Sodium cacodylate trihydrate pH 6.5 | 10% v/v 2-Propanol |
21 | 0.2 M KCl 0.005 M Magnesium chloride hexahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.5 | 0.9 M 1, 6-Hexanediol |
Natrix HT (HR2-131)
No | Salt | Buffer | Precipitant |
22 | 0.08 M Magnesium acetate tetrahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.5 | 15% v/v PEG 400 |
23 | 0.2 M KCl 0.01 M Magnesium chloride hexahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.5 | 10% w/v PEG 4000 |
24 | 0.2 M NH4CH3CO2 0.01 M Calcium chloride dihydrate | 0.05 M Sodium cacodylate trihydrate pH 6.5 | 10% w/v PEG 4000 |
25 | 0.08 M Magnesium acetate tetrahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.5 | 30% w/v PEG 4000 |
26 | 0.2 M KCl 0.1 M Magnesium acetate tetrahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.5 | 10% w/v PEG 8000 |
27 | 0.2 M NH4CH3CO2 0.01 M Mg(CH3COO)2 • 4H2O | 0.05 M Sodium cacodylate trihydrate pH 6.5 | 30% w/v PEG 8000 |
28 | 0.05 M Magnesium sulfate hydrate | 0.05 M HEPES Sodium pH 7.0 | 1.6 M Lithium sulfate monohydrate |
29 | 0.01 M Magnesium chloride hexahydrate | 0.05 M HEPES Sodium pH 7.0 | 4.0 M Lithium chloride |
30 | 0.01 M Magnesium chloride hexahydrate | 0.05 M HEPES Sodium pH 7.0 | 1.6 M (NH4)2SO4 |
31 | 0.005 M Magnesium chloride hexahydrate | 0.05 M HEPES Sodium pH 7.0 | 25% v/v PEG monomethyl ether 550 |
32 | 0.2 M KCl 0.01 M Magnesium chloride hexahydrate | 0.05 M HEPES Sodium pH 7.0 | 1.7 M 1, 6-Hexanediol |
33 | 0.2 M Ammonium chloride 0.01 M Magnesium chloride hexahydrate | 0.05 M HEPES pH 7.0 | 2.5 M 1, 6-Hexanediol |
34 | 0.1 M KCl 0.005 M Magnesium sulfate hydrate | 0.05 M HEPES Sodium pH 7.0 | 15% v/v MPD |
35 | 0.1 M KCl 0.01 M Magnesium chloride hexahydrate | 0.05 M HEPES Sodium pH 7.0 | 5% v/v PEG 400 |
36 | 0.1 M KCl 0.01 M Calcium chloride dihydrate | 0.05 M HEPES Sodium pH 7.0 | 10% v/v PEG 400 |
37 | 0.2 M KCl 0.025 M Magnesium sulfate hydrate | 0.05 M HEPES Sodium pH 7.0 | 20% v/v PEG 200 |
38 | 0.2 M Ammonium acetate 0.15 M Magnesium acetate tetrahydrate | 0.05 M HEPES Sodium pH 7.0 | 5% w/v PEG 4000 |
39 | 0.1 M Ammonium acetate 0.02 M Magnesium chloride hexahydrate | 0.05 M HEPES Sodium pH 7.0 | 5% w/v PEG 8000 |
40 | 0.01 M Magnesium chloride hexahydrate | 0.05 M TRIS hydrochloride pH 7.5 | 1.6 M (NH4)2SO4 |
41 | 0.1 M KCl 0.015 M MgCl2·6H2O | 0.05 M Tris-HCl pH 7.5 | 10% v/v PEG monomethyl ether 550 |
42 | 0.01 M MgCl2·6H2O | 0.05 M Tris-HCl pH 7.5 | 5% v/v 2-Propanol |
Natrix HT (HR2-131)
No | Salt | Buffer | Precipitant | Additive |
43 | 0.05 M Ammonium acetate 0.01 M MgCl2·6H2O | 0.05 M Tris-HCl pH 7.5 | 10% v/v MPD |
|
44 | 0.2 M KCl 0.05 M MgCl2·6H2O | 0.05 M Tris-HCl pH 7.5 | 10% w/v PEG 4000 |
|
45 | 0.025 M Magnesium sulfate hydrate | 0.05 M Tris-HCl pH 8.5 | 1.8 M Ammonium sulfate |
|
46 | 0.005 M Magnesium sulfate hydrate | 0.05 M Tris-HCl pH 8.5 | 2.9 M 1, 6-Hexanediol |
|
47 | 0.1 M KCl 0.01 M MgCl2·6H2O | 0.05 M Tris-HCl pH 8.5 | 30% v/v PEG 400 |
|
48 | 0.2 M Ammonium chloride 0.01 M CaCl2·2H2O | 0.05 M Tris-HCl pH 8.5 | 30% w/v PEG 4000 |
|
49 | 0.04 M Lithium chloride 0.02 M MgCl2·6H2O | 0.04 M Sodium cacodylate trihydrate pH 5.5 | 30% v/v MPD | 0.002 M Hexammine cobalt(III) chloride |
50 | 0.08 M NaCl 0.02 M MgCl2·6H2O | 0.04 M Sodium cacodylate trihydrate pH 5.5 | 35% v/v MPD | 0.002 M Hexammine cobalt(III) chloride |
51 | 0.012 M NaCl 0.08 M KCl | 0.04 M Sodium cacodylate trihydrate pH 5.5 | 45% v/v MPD | 0.002 M Hexammine cobalt(III) chloride |
52 | 0.02 M MgCl2 6H2O | 0.04 M Sodium cacodylate trihydrate pH 5.5 | 40% v/v MPD | 0.002 M Hexammine cobalt(III) chloride |
53 | 2 mM CaCl2·2H2O | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 1.8 M (NH4)2SO4 | 0.5 mM Spermine |
54 |
| 0.05 M Sodium cacodylate trihydrate pH 6.0 | 35% v/v Tacsimate pH 6.0 | 0.001 M Spermine |
55 | 0.1 M NaCl | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 10% w/v PEG 4000 | 0.0005 M Spermine |
56 | 0.05 M KCl | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 10% w/v PEG 8000 | 0.0005 M Spermine, 0.0005 M L-Argininamide dihydrochloride |
57 | 0.1 M KCl | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 16% w/v PEG 1000 | 0.0005 M Spermine |
58 | 5 mM MgCl2·6H2O 2 mM CaCl2·2H2O | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 15% v/v 2-Propano | 1 mM Spermine |
59 | 0.075 M NaCl 2 mM CaCl2·2H2O | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 30% w/v 1, 6-Hexanediol | 0. 5 mM Spermine |
60 | 0.02 M Magnesium sulfate hydrate 0.002 M Cobalt(II) chloride hexahydrate | 0.05 M Sodium cacodylate trihydrate pH 6.0 | 25% v/v MPD | 0.0005 M Spermine |
Natrix HT (HR2-131) | ||||
No | Salt | Buffer | Precipitant | Additive |
61 |
| 0.05 M Sodium cacodylate trihydrate pH 6.0 | 30% v/v MPD |
|
62 | 0.08 M NaCl 0.012 M KCl 0.02 M MgCl2·6H2O | 0.04 M Sodium cacodylate trihydrate pH 6.0 | 30% v/v MPD | 0.012 M Spermine tetrahydrochloride |
63 | 0.08 M NaCl 0.02 M MgCl2·6H2O | 0.04 M Sodium cacodylate trihydrate pH 6.0 | 35% v/v MPD | 0.012 M Spermine tetrahydrochloride |
64 | 0.08 M Strontium chloride hexahydrate | 0.04 M Sodium cacodylate trihydrate pH 6.0 | 35% v/v MPD | 0.012 M Spermine tetrahydrochloride |
65 | 0.08 M KCl 0.02 M Barium chloride dihydrate | 0.04 M Sodium cacodylate trihydrate pH 6.0 | 40% v/v MPD | 0.012 M Spermine tetrahydrochloride |
66 | 0.08 M KCl 0.02 M MgCl2·6H2O | 0.04 M Sodium cacodylate trihydrate pH 6.0 | 45% v/v MPD | 0.012 M Spermine tetrahydrochloride |
67 | 0.08 M NaCl | 0.04 M Sodium cacodylate trihydrate pH 6.0 | 45% v/v MPD | 0.012 M Spermine tetrahydrochloride |
68 | 0.08 M NaCl 0.02 M Barium chloride dihydrate | 0.04 M Sodium cacodylate trihydrate pH 6.0 | 45% v/v MPD | 0.012 M Spermine tetrahydrochloride |
69 | 0.012 M NaCl 0.08 M KCl | 0.04 M Sodium cacodylate trihydrate pH 6.0 | 50% v/v MPD | 0.012 M Spermine tetrahydrochloride |
70 | 0.08 M KCl | 0.04 M Sodium cacodylate trihydrate pH 6.0 | 55% v/v MPD | 0.012 M Spermine tetrahydrochloride |
71 | 0.018 M MgCl2·6H2O | 0.05 M Sodium cacodylate trihydrate pH 6.5 | 10% v/v 2-Propanol | 0.003 M Spermine |
72 | 0.02 M MgCl2·6H2O | 0.05 M MOPS pH 7.0 | 2.0 M Ammonium sulfate | 0.0005 M Spermine |
73 |
| 0.05 M HEPES sodium pH 7.0 | 40% v/v Tacsimate pH 7.0 | 2 mM Spermine 2 mM Hexammine cobalt(III) chloride |
74 | 0.02 M MgCl2·6H2O | 0.05 M MOPS pH 7.0 | 55% v/v Tacsimate pH 7.0 | 0.002 M Hexammine cobalt(III) chloride |
75 | 0.02 M MgCl2·6H2O | 0.05 M Sodium cacodylate trihydrate pH 7.0 | 15% v/v 2-Propanol | 0.001 M Hexammine cobalt(III) chloride, 0.001 M Spermine |
76 | 0.005 M MgCl2·6H2O | 0.05 M MOPS pH 7.0 | 25% v/v 1, 4-Dioxane | 0.001 M Spermine |
77 | 0.01 M MgCl2·6H2O, 0.002 M Barium chloride dihydrate | 0.05 M MOPS pH 7.0 | 30% v/v 1, 4-Dioxane |
|
78 | 1 mM MgCl2·6H2O 0.002 M CaCl2·2H2O | 0.05 M MOPS pH 7.0 | 15% v/v MPD |
|
79 | 0.08 M Strontium chloride hexahydrate 0.02 M MgCl2·6H2O | 0.04 M Sodium cacodylate trihydrate pH 7.0 | 20% v/v MPD | 0.012 M Spermine tetrahydrochloride |
80 | 0.08 M NaCl | 0.04 M Sodium cacodylate trihydrate pH 7.0 | 30% v/v MPD | 0.012 M Spermine tetrahydrochloride |
81 | 0.04 M Lithium chloride 0.08 M Strontium chloride hexahydrate | 0.04 M Sodium cacodylate trihydrate pH 7.0 | 30% v/v MPD | 0.012 M Spermine tetrahydrochloride |
82 | 0.04 M Lithium chloride, 0.08 M Strontium chloride hexahydrate, 0.02 M MgCl2·6H2O | 0.04 M Sodium cacodylate trihydrate pH 7.0 | 30% v/v MPD | 0.012 M Spermine tetrahydrochloride |
83 | 0.08 M NaCl 0.012 M KCl 0.02 M MgCl2·6H2O | 0.04 M Sodium cacodylate trihydrate pH 7.0 | 35% v/v MPD | 0.012 M Spermine tetrahydrochloride |
84 | 0.012 M NaCl 0.08 M KCl | 0.04 M Sodium cacodylate trihydrate pH 7.0 | 40% v/v MPD | 0.012 M Spermine tetrahydrochloride |
85 | 0.08 M NaCl 0.02 M Barium chloride dihydrate | 0.04 M Sodium cacodylate trihydrate pH 7.0 | 40% v/v MPD | 0.012 M Spermine tetrahydrochloride |
86 | 0.08 M NaCl 0.02 M MgCl2·6H2O | 0.04 M Sodium cacodylate trihydrate pH 7.0 | 40% v/v MPD | 0.012 M Spermine tetrahydrochloride |
87 | 0.08 M KCl, 0.02 M Barium chloride dihydrate | 0.04 M Sodium cacodylate trihydrate pH 7.0 | 40% v/v MPD | 0.012 M Spermine tetrahydrochloride |
88 | 0.08 M KCl 0.02 M MgCl2·6H2O | 0.04 M Sodium cacodylate trihydrate pH 7.0 | 50% v/v MPD | 0.012 M Spermine tetrahydrochloride |
89 | 0.08 M KCl | 0.04 M Sodium cacodylate trihydrate pH 7.0 | 60% v/v MPD | 0.012 M Spermine tetrahydrochloride |
90 | 0.02 M MgCl2·6H2O 0.002 M Cobalt(II) chloride hexahydrate | 0.05 M HEPES sodium pH 7.5 | 2.0 M (NH4)2SO4 | 1 mM Spermine |
91 | 0.02 M MgCl2·6H2O | 0.05 M PIPES pH 7.5 | 4% w/v PEG 8000 | 0.001 M Spermine |
92 | 0.015 M MgCl2·6H2O, 2 mM Barium chloride dihydrate | 0.05 M PIPES pH 7.5 | 7% v/v 2-Propanol | 0. 5 mM Spermine |
93 | 0.02 M MgCl2·6H2O | 0.05 M PIPES pH 7.5 | 10% w/v 1, 6-Hexanediol | 0.001 M Spermine |
94 | 0.01 M MgCl2·6H2O | 0.05 M HEPES sodium pH 7.5 | 15% v/v MPD | 0.0015 M Spermine |
95 | 0.2 M CaCl2·2H2O | 0.05 M HEPES sodium pH 7.5 | 28% v/v PEG 400 | 0.002 M Spermine |
96 | 2 mM Copper(II) chloride dihydrate | 0.05 M Tris-HCl pH 8.5 | 1.8 M Lithium sulfate monohydrate | 0.5 mM Spermine |
SaltRx HT (HR2-136)
No | Salt | Buffer |
1 | 1.8 M Sodium acetate trihydrate pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
2 | 2.8 M Sodium acetate trihydrate pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
3 | 1.5 M Ammonium chloride | 0.1 M Sodium acetate trihydrate pH 4.6 |
4 | 1.5 M Ammonium chloride | 0.1 M BIS-TRIS propane pH 7.0 |
5 | 1.5 M Ammonium chloride | 0.1 M Tris pH 8.5 |
6 | 3.5 M Ammonium chloride | 0.1 M Sodium acetate trihydrate pH 4.6 |
7 | 3.5 M Ammonium chloride | 0.1 M BIS-TRIS propane pH 7.0 |
8 | 3.5 M Ammonium chloride | 0.1 M Tris pH 8.5 |
9 | 2.2 M NaCl | 0.1 M Sodium acetate trihydrate pH 4.6 |
10 | 2.2 M NaCl | 0.1 M BIS-TRIS propane pH 7.0 |
11 | 2.2 M NaCl | 0.1 M Tris pH 8.5 |
12 | 3.2 M NaCl | 0.1 M Sodium acetate trihydrate pH 4.6 |
13 | 3.2 M NaCl | 0.1 M BIS-TRIS propane pH 7.0 |
14 | 3.2 M NaCl | 0.1 M Tris pH 8.5 |
15 | 1.0 M Ammonium citrate dibasic | 0.1 M Sodium acetate trihydrate pH 4.6 |
16 | 1.8 M Ammonium citrate dibasic | 0.1 M Sodium acetate trihydrate pH 4.6 |
17 | 1.0 M Ammonium citrate tribasic pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
18 | 2.0 M Ammonium citrate tribasic pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
19 | 0.7 M Sodium citrate tribasic dihydrate | 0.1 M BIS-TRIS propane pH 7.0 |
20 | 0.7 M Sodium citrate tribasic dihydrate | 0.1 M Tris pH 8.5 |
21 | 1.2 M Sodium citrate tribasic dihydrate | 0.1 M BIS-TRIS propane pH 7.0 |
22 | 1.2 M Sodium citrate tribasic dihydrate | 0.1 M Tris pH 8.5 |
23 | 0.4 M Magnesium formate dihydrate | 0.1 M Sodium acetate trihydrate pH 4.6 |
24 | 0.4 M Magnesium formate dihydrate | 0.1 M BIS-TRIS propane pH 7.0 |
25 | 0.4 M Magnesium formate dihydrate | 0.1 M Tris pH 8.5 |
26 | 0.7 M Magnesium formate dihydrate | 0.1 M BIS-TRIS propane pH 7.0 |
27 | 2.0 M Sodium formate | 0.1 M Sodium acetate trihydrate pH 4.6 |
28 | 2.0 M Sodium formate | 0.1 M BIS-TRIS propane pH 7.0 |
29 | 2.0 M Sodium formate | 0.1 M Tris pH 8.5 |
30 | 3.5 M Sodium formate | 0.1 M Sodium acetate trihydrate pH 4.6 |
31 | 3.5 M Sodium formate | 0.1 M BIS-TRIS propane pH 7.0 |
32 | 3.5 M Sodium formate | 0.1 M Tris pH 8.5 |
33 | 1.2 M DL-Malic acid pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
34 | 2.2 M DL-Malic acid pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
35 | 1.4 M Sodium malonate pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
36 | 2.4 M Sodium malonate pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
37 | 2.5 M Ammonium nitrate | 0.1 M Sodium acetate trihydrate pH 4.6 |
38 | 2.5 M Ammonium nitrate | 0.1 M BIS-TRIS propane pH 7.0 |
39 | 2.5 M Ammonium nitrate | 0.1 M Tris pH 8.5 |
40 | 6.0 M Ammonium nitrate | 0.1 M Sodium acetate trihydrate pH 4.6 |
41 | 6.0 M Ammonium nitrate | 0.1 M BIS-TRIS propane pH 7.0 |
42 | 6.0 M Ammonium nitrate | 0.1 M Tris pH 8.5 |
43 | 1.5 M Sodium nitrate | 0.1 M Sodium acetate trihydrate pH 4.6 |
44 | 1.5 M Sodium nitrate | 0.1 M BIS-TRIS propane pH 7.0 |
45 | 1.5 M Sodium nitrate | 0.1 M Tris pH 8.5 |
46 | 4.0 M Sodium nitrate | 0.1 M Sodium acetate trihydrate pH 4.6 |
47 | 4.0 M Sodium nitrate | 0.1 M BIS-TRIS propane pH 7.0 |
48 | 4.0 M Sodium nitrate | 0.1 M Tris pH 8.5 |
49 | 1.0 M Ammonium phosphate monobasic | 0.1 M Sodium acetate trihydrate pH 4.6 |
50 | 1.8 M Ammonium phosphate monobasic | 0.1 M Sodium acetate trihydrate pH 4.6 |
51 | 1.5 M Ammonium phosphate dibasic | 0.1 M Tris pH 8.5 |
52 | 2.4 M Ammonium phosphate dibasic | 0.1 M Tris pH 8.5 |
53 | 1.0 M Sodium phosphate monobasic monohydrate | Potassium phosphate dibasic / pH 5.0 |
54 | 1.0 M Sodium phosphate monobasic monohydrate | Potassium phosphate dibasic / pH 6.9 |
55 | 1.0 M Sodium phosphate monobasic monohydrate | Potassium phosphate dibasic / pH 8.2 |
56 | 1.8 M Sodium phosphate monobasic monohydrate | Potassium phosphate dibasic / pH 5.0 |
57 | 1.8 M Sodium phosphate monobasic monohydrate | Potassium phosphate dibasic / pH 6.9 |
58 | 1.8 M Sodium phosphate monobasic monohydrate | Potassium phosphate dibasic / pH 8.2 |
59 | 0.5 M Succinic acid pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
60 | 1.0 M Succinic acid pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
61 | 1.5 M Ammonium sulfate | 0.1 M Sodium acetate trihydrate pH 4.6 |
62 | 1.5 M Ammonium sulfate | 0.1 M BIS-TRIS propane pH 7.0 |
63 | 1.5 M Ammonium sulfate | 0.1 M Tris pH 8.5 |
64 | 2.5 M Ammonium sulfate | 0.1 M Sodium acetate trihydrate pH 4.6 |
65 | 2.5 M Ammonium sulfate | 0.1 M BIS-TRIS propane pH 7.0 |
66 | 2.5 M Ammonium sulfate | 0.1 M Tris pH 8.5 |
67 | 0.8 M Lithium sulfate monohydrate | 0.1 M Sodium acetate trihydrate pH 4.6 |
68 | 0.8 M Lithium sulfate monohydrate | 0.1 M BIS-TRIS propane pH 7.0 |
69 | 0.8 M Lithium sulfate monohydrate | 0.1 M Tris pH 8.5 |
70 | 1.5 M Lithium sulfate monohydrate | 0.1 M Sodium acetate trihydrate pH 4.6 |
71 | 1.5 M Lithium sulfate monohydrate | 0.1 M BIS-TRIS propane pH 7.0 |
72 | 1.5 M Lithium sulfate monohydrate | 0.1 M Tris pH 8.5 |
73 | 1.0 M Magnesium sulfate hydrate | 0.1 M Sodium acetate trihydrate pH 4.6 |
74 | 1.0 M Magnesium sulfate hydrate | 0.1 M BIS-TRIS propane pH 7.0 |
75 | 1.0 M Magnesium sulfate hydrate | 0.1 M Tris pH 8.5 |
76 | 1.8 M Magnesium sulfate hydrate | 0.1 M Sodium acetate trihydrate pH 4.6 |
77 | 1.8 M Magnesium sulfate hydrate | 0.1 M BIS-TRIS propane pH 7.0 |
78 | 1.8 M Magnesium sulfate hydrate | 0.1 M Tris pH 8.5 |
79 | 0.7 M Ammonium tartrate dibasic | 0.1 M Sodium acetate trihydrate pH 4.6 |
80 | 0.7 M Ammonium tartrate dibasic | 0.1 M BIS-TRIS propane pH 7.0 |
81 | 0.7 M Ammonium tartrate dibasic | 0.1 M Tris pH 8.5 |
82 | 1.0 M Ammonium tartrate dibasic | 0.1 M Sodium acetate trihydrate pH 4.6 |
83 | 1.3 M Ammonium tartrate dibasic | 0.1 M BIS-TRIS propane pH 7.0 |
84 | 1.4 M Ammonium tartrate dibasic | 0.1 M Tris pH 8.5 |
85 | 0.6 M Potassium sodium tartrate tetrahydrate | 0.1 M BIS-TRIS propane pH 7.0 |
86 | 1.2 M Potassium sodium tartrate tetrahydrate | 0.1 M BIS-TRIS propane pH 7.0 |
87 | 0.6 M Potassium sodium tartrate tetrahydrate | 0.1 M Tris pH 8.5 |
88 | 1.2 M Potassium sodium tartrate tetrahydrate | 0.1 M Tris pH 8.5 |
89 | 0.5 M Potassium thiocyanate | 0.1 M Sodium acetate trihydrate pH 4.6 |
90 | 0.5 M Potassium thiocyanate | 0.1 M BIS-TRIS propane pH 7.0 |
91 | 0.5 M Potassium thiocyanate | 0.1 M Tris pH 8.5 |
92 | 4.0 M Ammonium acetate | 0.1 M Sodium acetate trihydrate pH 4.6 |
93 | 4.0 M Ammonium acetate | 0.1 M BIS-TRIS propane pH 7.0 |
94 | 4.0 M Ammonium acetate | 0.1 M Tris pH 8.5 |
95 | 35% v/v Tacsimate pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
96 | 60% v/v Tacsimate pH 7.0 | 0.1 M BIS-TRIS propane pH 7.0 |
PEGRx HT (HR2-086)
No | Buffer | Polymer |
1 | 0.1 M Citric acid pH 3.5 | 34% v/v PEG 200 |
2 | 0.1 M Sodium citrate tribasic dihydrate pH 5.5 | 38% v/v PEG 200 |
3 | 0.1 M HEPES pH 7.5 | 42% v/v PEG 200 |
4 | 0.1 M Sodium acetate trihydrate pH 4.5 | 30% v/v PEG 300 |
5 | 0.1 M BIS-TRIS pH 6.5 | 25% v/v PEG 300 |
6 | 0.1 M Bicine pH 8.5 | 20% v/v PEG 300 |
7 | 0.1 M Sodium acetate trihydrate pH 4.0 | 15% v/v PEG 400 |
8 | 0.1 M MES monohydrate pH 6.0 | 22% v/v PEG 400 |
9 | 0.1 M Tris pH 8.0 | 30% v/v PEG 400 |
10 | 0.1 M Sodium citrate tribasic dihydrate pH 5.0 | 30% v/v PEG MME 550 |
11 | 0.1 M Imidazole pH 7.0 | 25% v/v PEG MME 550 |
12 | 0.1 M BIS-TRIS propane pH 9.0 | 20% v/v PEG MME 550 |
13 | 0.1 M Sodium acetate trihydrate pH 4.0 | 10% v/v Jeffamine® M-600® pH 7.0 |
14 | 0.1 M MES monohydrate pH 6.0 | 20% v/v Jeffamine® M-600® pH 7.0 |
15 | 0.1 M Tris pH 8.0 | 30% v/v Jeffamine® M-600® pH 7.0 |
16 | 0.1 M Citric acid pH 3.5 | 14% w/v PEG 1000 |
17 | 0.1 M Sodium citrate tribasic dihydrate pH 5.5 | 22% w/v PEG 1000 |
18 | 0.1 M HEPES pH 7.5 | 30% w/v PEG 1000 |
19 | 0.1 M Sodium acetate trihydrate pH 4.5 | 30% w/v PEG 1500 |
20 | 0.1 M BIS-TRIS pH 6.5 | 20% w/v PEG 1500 |
21 | 0.1 M Bicine pH 8.5 | 15% w/v PEG 1500 |
22 | 0.1 M Sodium acetate trihydrate pH 4.0 | 10% w/v PEG MME 2000 |
23 | 0.1 M MES monohydrate pH 6.0 | 20% w/v PEG MME 2000 |
24 | 0.1 M Tris pH 8.0 | 30% w/v PEG MME 2000 |
25 | 0.1 M Sodium citrate tribasic dihydrate pH 5.0 | 30% v/v Jeffamine® ED-2001 pH 7.0 |
26 | 0.1 M Imidazole pH 7.0 | 20% v/v Jeffamine® ED-2001 pH 7.0 |
27 | 0.1 M BIS-TRIS propane pH 9.0 | 10% v/v Jeffamine® ED-2001 pH 7.0 |
28 | 0.1 M Citric acid pH 3.5 | 25% w/v PEG 3350 |
29 | 0.1 M Sodium citrate tribasic dihydrate pH 5.5 | 18% w/v PEG 3350 |
30 | 0.1 M HEPES pH 7.5 | 12% w/v PEG 3350 |
31 | 0.1 M Sodium acetate trihydrate pH 4.0 | 10% w/v PEG 4000 |
32 | 0.1 M MES monohydrate pH 6.0 | 14% w/v PEG 4000 |
33 | 0.1 M Tris pH 8.0 | 28% w/v PEG 4000 |
34 | 0.1 M Sodium acetate trihydrate pH 4.5 | 30% PEG MME 5000 |
35 | 0.1 M BIS-TRIS pH 6.5 | 20% w/v PEG MME 5000 |
36 | 0.1 M Bicine pH 8.5 | 8% w/v PEG MME 5000 |
37 | 0.1 M Sodium citrate tribasic dihydrate pH 5.0 | 10% w/v PEG 6000 |
38 | 0.1 M Imidazole pH 7.0 | 20% w/v PEG 6000 |
39 | 0.1 M BIS-TRIS propane pH 9.0 | 30% w/v PEG 6000 |
40 | 0.1 M Citric acid pH 3.5 | 28% w/v PEG 8000 |
41 | 0.1 M Sodium citrate tribasic dihydrate pH 5.5 | 16% w/v PEG 8000 |
42 | 0.1 M HEPES pH 7.5 | 4% w/v PEG 8000 |
43 | 0.1 M Sodium acetate trihydrate pH 4.5 | 10% w/v PEG 10,000 |
44 | 0.1 M BIS-TRIS pH 6.5 | 16% w/v PEG 10,000 |
45 | 0.1 M Bicine pH 8.5 | 20% w/v PEG 10,000 |
46 | 0.1 M Sodium citrate tribasic dihydrate pH 5.0 | 18% w/v PEG 20,000 |
47 | 0.1 M Imidazole pH 7.0 | 12% w/v PEG 20,000 |
48 | 0.1 M BIS-TRIS propane pH 9.0, | 8% w/v PEG 20,000 |
No | Buffer | Buffer | Polymer |
49 | 0.8 M Lithium sulfate monohydrate | 0.1 M Sodium acetate trihydrate pH 4.0 | 4% v/v PEG 200 |
50 | 0.2 M Lithium sulfate monohydrate | 0.1 M Sodium citrate tribasic dihydrate pH 5.0 | 26% v/v PEG 200 |
51 | 0.05 M CaCl2·2H2O | 0.1 M MES monohydrate pH 6.0 | 45% v/v PEG 200 |
52 | 28% v/v 2-Propanol | 0.1 M BIS-TRIS pH 6.5 | 3% v/v PEG 200 |
53 | 20% v/v Tacsimate pH 7.0 | 0.1 M HEPES pH 7.5 | 2% v/v PEG 200 |
54 | 10% v/v 2-Propanol | 0.1 M Sodium citrate tribasic dihydrate pH 5.0 | 26% v/v PEG 400 |
55 | 0.2 M Ammonium acetate | 0.1 M Sodium citrate tribasic dihydrate pH 5.5 | 24% v/v PEG 400 |
56 | 0.2 M Ammonium sulfate | 0.1 M BIS-TRIS pH 6.5 | 18% v/v PEG 400 |
57 | 0.19 mM CYMAL®-7 | 0.1 M HEPES pH 7.5 | 40% v/v PEG 400 |
58 | 6% v/v 2-Propanol | 0.1 M Sodium acetate trihydrate pH 4.5 | 26% v/v PEG MME 550 |
59 | 1.8 M Ammonium sulfate | 0.1 M BIS-TRIS pH 6.5 | 2% v/v PEG MME 550 |
60 | 0.15 M DL-Malic acid pH 7.0 | 0.1 M Imidazole pH 7.0 | 22% v/v PEG MME 550 |
61 | 0.1 M Succinic acid pH 7.0 | 0.1 M Bicine pH 8.5 | 30% v/v PEG MME 550 |
62 | 0.1 M Lithium sulfate monohydrate | 0.1 M Sodium citrate tribasic dihydrate pH 5.5 | 20% w/v PEG 1000 |
63 | 0.1 M Sodium malonate pH 8.0 | 0.1 M Tris pH 8.0 | 30% w/v PEG 1000 |
64 | 4% v/v MPD | 0.1 M Citric acid pH 3.5 | 20% w/v PEG 1500 |
65 | 0.2 M L-Proline | 0.1 M HEPES pH 7.5 | 24% w/v PEG 1500 |
66 | 10% v/v 2-Propanol | 0.1 M Bicine pH 8.5 | 30% w/v PEG 1500 |
67 | 0.1 M NaCl | 0.1 M BIS-TRIS propane pH 9.0 | 25% w/v PEG 1500 |
68 | 0.02 M Nickel(II) chloride hexahydrate 0.02 M MgCl2·6H2O 0.02 M Cadmium chloride hydrate | 0.1 M Sodium acetate trihydrate pH 4.5 | 24% w/v PEG MME 2000 |
69 | 20% v/v 2-Propanol | 0.1 M MES monohydrate pH 6.0 | 20% w/v PEG MME 2000 |
70 | 0.2 M Ammonium citrate tribasic pH 7.0 | 0.1 M Imidazole pH 7.0 | 20% w/v PEG MME 2000 |
71 | 4.0 M Potassium formate | 0.1 M BIS-TRIS propane pH 9.0 | 2% w/v PEG MME 2000 |
72 | 50% v/v Tacsimate pH 4.0 | 0.1 M Sodium acetate trihydrate pH 4.5 | 1% w/v PEG 3350 |
73 | 0.10% w/v n-Octyl-β-D-glucoside | 0.1 M Sodium citrate tribasic dihydrate pH 5.5 | 22% w/v PEG 3350 |
74 | 2% v/v Tacsimate pH 7.0 5% v/v 2-Propanol | 0.1 M Imidazole pH 7.0 | 8% w/v PEG 3350 |
75 | 2% v/v 1, 4-Dioxane | 0.1 M Tris pH 8.0 | 15% w/v PEG 3350 |
76 | 18% v/v 2-Propanol | 0.1 M Sodium citrate tribasic dihydrate pH 5.5 | 20% w/v PEG 4000 |
77 | 6% v/v Tacsimate pH 6.0 | 0.1 M MES monohydrate pH 6.0 | 25% w/v PEG 4000 |
78 | 0.2 M Magnesium formate dihydrate | 0.1 M Sodium acetate trihydrate pH 4.0 | 18% w/v PEG MME 5000 |
79 | 2% v/v PEG 400 | 0.1 M Imidazole pH 7.0 | 24% w/v PEG MME 5000 |
80 | 0.2 M Sodium formate | 0.1 M Bicine pH 8.5 | 20% w/v PEG MME 5000 |
81 | 4% v/v 2-Propanol | 0.1 M BIS-TRIS propane pH 9.0 | 20% w/v PEG MME 5000 |
82 | 6% v/v Ethylene glycol | 0.1 M Citric acid pH 3.5 | 10% w/v PEG 6000 |
83 | 0.15 M Lithium sulfate monohydrate | 0.1 M Citric acid pH 3.5 | 18% PEG 6000 |
84 | 10% v/v 2-Propanol | 0.1 M Sodium acetate trihydrate pH 4.0 | 22% w/v PEG 6000 |
85 | 0.2 M NaCl | 0.1 M Sodium acetate trihydrate pH 4.0 | 22% w/v PEG 8000 |
86 | 20% v/v 2-Propanol | 0.1 M Tris pH 8.0 | 5% w/v PEG 8000 |
87 | 10% v/v PEG 200 | 0.1 M BIS-TRIS propane pH 9.0 | 18% w/v PEG 8000 |
88 | 15% v/v 2-Propanol | 0.1 M Sodium citrate tribasic dihydrate pH 5.0 | 10% w/v PEG 10000 |
89 | 0.4 M Sodium malonate pH 6.0 | 0.1 M MES monohydrate pH 6.0 | 0.5% w/v PEG 10000 |
90 | 0.2 M Potassium sodium tartrate tetrahydrate | 0.1 M BIS-TRIS pH 6.5 | 10% w/v PEG 10000 |
91 | 5% v/v MPD | 0.1 M HEPES pH 7.5 | 10% w/v PEG 10000 |
92 | 0.2 M Ammonium acetate | 0.1 M Tris pH 8.0 | 16% w/v PEG 10000 |
93 | 5% v/v 2-Propanol | 0.1 M Citric acid pH 3.5 | 6% w/v PEG 20000 |
94 | 1.0 M Sodium malonate pH 5.0 | 0.1 M Sodium acetate trihydrate pH 4.5 | 2% w/v PEG 20000 |
95 | 0.2 M MgCl2·6H2O | 0.1 M Sodium citrate tribasic dihydrate pH 5.0 | 10% w/v PEG 20000 |
96 | 3% w/v Dextran sulfate sodium salt | 0.1 M Bicine pH 8.5 | 15% w/v PEG 20000 |
Crystal Screen HT (HR2-130)
No | Salt | Buffer | Precipitant |
1 | 0.02 M CaCl2·2H2O | 0.1 M Sodium acetate trihydrate pH 4.6 | 30% v/v MPD |
2 | 0.4 M Potassium sodium tartrate tetrahydrate |
|
|
3 | 0.4 M Ammonium phosphate monobasic |
|
|
4 | 0.1 M Tris-HCl pH 8.5 | 2.0 M Ammonium sulfate |
|
5 | 0.2 M Sodium citrate tribasic dihydrate | 0.1 M HEPES sodium pH 7.5 | 30% v/v MPD |
6 | 0.2 M MgCl2·6H2O | 0.1 M Tris-HCl pH 8.5 | 30% w/v PEG 4000 |
7 | 0.1 M Sodium cacodylate trihydrate pH 6.5 | 1.4 M Sodium acetate trihydrate |
|
8 | 0.2 M Sodium citrate tribasic dihydrate | 0.1 M Sodium cacodylate trihydrate pH 6.5 | 30% v/v 2-Propanol |
9 | 0.2 M Ammonium acetate | 0.1 M Sodium citrate tribasic dihydrate pH 5.6 | 30% w/v PEG 4000 |
10 | 0.2 M Ammonium acetate | 0.1 M Sodium acetate trihydrate pH 4.6 | 30% w/v PEG 4000 |
11 | 0.1 M Sodium citrate tribasic dihydrate pH 5.6 | 1.0 M Ammonium phosphate monobasic |
|
12 | 0.2 M MgCl2·6H2O | 0.1 M HEPES sodium pH 7.5 | 30% v/v 2-Propanol |
13 | 0.2 M Sodium citrate tribasic dihydrate | 0.1 M Tris-HCl pH 8.5 | 30% v/v PEG 400 |
14 | 0.2 M CaCl2·2H2O | 0.1 M HEPES sodium pH 7.5 | 28% v/v PEG 400 |
15 | 0.2 M Ammonium sulfate | 0.1 M Sodium cacodylate trihydrate pH 6.5 | 30% w/v PEG 8000 |
16 | 0.1 M HEPES sodium pH 7.5 | 1.5 M Lithium sulfate monohydrate |
|
17 | 0.2 M Lithium sulfate monohydrate | 0.1 M Tris-HCl pH 8.5 | 30% w/v PEG 4000 |
18 | 0.2 M Magnesium acetate tetrahydrate | 0.1 M Sodium cacodylate trihydrate pH 6.5 | 20% w/v PEG 8000 |
19 | 0.2 M Ammonium acetate | 0.1 M Tris-HCl pH 8.5 | 30% v/v 2-Propanol |
20 | 0.2 M Ammonium sulfate | 0.1 M Sodium acetate trihydrate pH 4.6 | 25% w/v PEG 4000 |
21 | 0.2 M Magnesium acetate tetrahydrate | 0.1 M Sodium cacodylate trihydrate pH 6.5 | 30% v/v MPD |
22 | 0.2 M Sodium acetate trihydrate | 0.1 M Tris-HCl pH 8.5 | 30% w/v PEG 4000 |
23 | 0.2 M MgCl2·6H2O | 0.1 M HEPES sodium pH 7.5 | 30% v/v PEG 400 |
24 | 0.2 M CaCl2·2H2O | 0.1 M Sodium acetate trihydrate pH 4.6 | 20% v/v 2-Propanol |
25 | 0.1 M Imidazole pH 6.5 | 1.0 M Sodium acetate trihydrate |
|
26 | 0.2 M Ammonium acetate | 0.1 M Sodium citrate tribasic dihydrate pH 5.6 | 30% v/v MPD |
27 | 0.2 M Sodium citrate tribasic dihydrate | 0.1 M HEPES sodium pH 7.5 | 20% v/v 2-Propanol |
28 | 0.2 M Sodium acetate trihydrate | 0.1 M Sodium cacodylate trihydrate pH 6.5 | 30% w/v PEG 8000 |
29 | 0.1 M HEPES sodium pH 7.5 | 0.8 M Potassium sodium tartrate tetrahydrate |
|
30 | 0.2 M Ammonium sulfate | 30% w/v PEG 8000 |
|
31 | 0.2 M Ammonium sulfate | 30% w/v PEG 4000 |
|
32 | 2.0 M Ammonium sulfate |
|
|
33 | 4.0 M Sodium formate |
|
|
34 | 0.1 M Sodium acetate trihydrate pH 4.6 | 2.0 M Sodium formate |
|
35 | 0.1 M HEPES sodium pH 7.5 | 0.8 M Sodium phosphate monobasic monohydrate | 0.8 M Potassium phosphate monobasic |
36 | 0.1 M Tris-HCl pH 8.5 | 8% w/v PEG 8000 |
|
37 | 0.1 M Sodium acetate trihydrate pH 4.6 | 8% w/v PEG 4000 |
|
38 | 0.1 M HEPES sodium pH 7.5 | 1.4 M Sodium citrate tribasic dihydrate |
|
|
|
|
|
39 | 0.1 M HEPES sodium pH 7.5 | 2% v/v PEG 400 | 2.0 M Ammonium sulfate |
40 | 0.1 M Sodium citrate tribasic dihydrate pH 5.6 | 20% v/v 2-Propanol | 20% w/v PEG 4000 |
41 | 0.1 M HEPES sodium pH 7.5 | 10% v/v 2-Propanol | 20% w/v PEG 4000 |
42 | 0.05 M Potassium phosphate monobasic | 20% w/v PEG 8000 |
|
43 | 30% w/v PEG 1500 |
|
|
44 | 0.2 M Magnesium formate dihydrate |
|
|
45 | 0.2 M Zinc acetate dihydrate | 0.1 M Sodium cacodylate trihydrate pH 6.5 | 18% w/v PEG 8000 |
46 | 0.2 M Calcium acetate hydrate | 0.1 M Sodium cacodylate trihydrate pH 6.5 | 18% w/v PEG 8000 |
47 | 0.1 M Sodium acetate trihydrate pH 4.6 | 2.0 M Ammonium sulfate |
|
48 | 0.1 M Tris-HCl pH 8.5 | 2.0 M Ammonium phosphate monobasic |
|
49 | 2.0 M NaCl | 10% w/v PEG 6000 |
|
50 | 0.5 M NaCl | 0.01 M MgCl2·6H2O | 0.01 M Hexadecyltrimethylammonium bromide |
51 | 25% v/v Ethylene glycol |
|
|
52 | 35% v/v 1 | 4-Dioxane |
|
53 | 2.0 M Ammonium sulfate | 5% v/v 2-Propanol |
|
54 | 1.0 M Imidazole pH 7.0 |
|
|
55 | 10% w/v PEG 1000 | 10% w/v PEG 8000 |
|
56 | 1.5 M NaCl | 10% v/v Ethanol |
|
57 | 0.1 M Sodium acetate trihydrate pH 4.6 | 2.0 M NaCl |
|
58 | 0.2 M NaCl | 0.1 M Sodium acetate trihydrate pH 4.6 | 30% v/v MPD |
59 | 0.01 M Cobalt(II) chloride hexahydrate | 0.1 M Sodium acetate trihydrate pH 4.6 | 1.0 M 1, 6-Hexanediol |
60 | 0.1 M Cadmium chloride hydrate | 0.1 M Sodium acetate trihydrate pH 4.6 | 30% v/v PEG 400 |
61 | 0.2 M Ammonium sulfate | 0.1 M Sodium acetate trihydrate pH 4.6 | 30% w/v PEG MME 2000 |
62 | 0.2 M Potassium sodium tartrate tetrahydrate | 0.1 M Sodium citrate tribasic dihydrate pH 5.6 | 2.0 M (NH4)2SO4 |
63 | 0.5 M (NH4)2SO4 | 0.1 M Sodium citrate tribasic dihydrate pH 5.6 | 1.0 M Lithium sulfate monohydrate |
64 | 0.5 M NaCl | 0.1 M Sodium citrate tribasic dihydrate pH 5.6 | 2% v/v Ethylene imine polymer |
65 | 0.1 M Sodium citrate tribasic dihydrate pH 5.6 | 35% v/v tert-Butanol |
|
66 | 0.01 M Iron(III) chloride hexahydrate | 0.1 M Sodium citrate tribasic dihydrate pH 5.6 | 10% v/v Jeffamine M-600 |
67 | 0.1 M Sodium citrate tribasic dihydrate pH 5.6 | 2.5 M 1 | 6-Hexanediol |
68 | 0.1 M MES monohydrate pH 6.5 | 1.6 M Magnesium sulfate heptahydrate |
|
69 | 0.1 M Sodium phosphate monobasic monohydrate, 0.1 M Potassium phosphate monobasic | 0.1 M MES monohydrate pH 6.5 | 2.0 M NaCl |
70 | 0.1 M MES monohydrate pH 6.5 | 12% w/v PEG 20000 |
|
71 | 1.6 M Ammonium sulfate | 0.1 M MES monohydrate pH 6.5 | 10% v/v 1, 4-Dioxane |
72 | 0.05 M Cesium chloride | 0.1 M MES monohydrate pH 6.5 | 30% v/v Jeffamine M-600 |
73 | 0.01 M Cobalt(II) chloride hexahydrate | 0.1 M MES monohydrate pH 6.5 | 1.8 M (NH4)2SO4 |
74 | 0.2 M Ammonium sulfate | 0.1 M MES monohydrate pH 6.5 | 30% w/v PEG MME 5000 |
75 | 0.1 M MES monohydrate pH 6.5 | 0.01 M Zinc sulfate heptahydrate | 25% v/v PEG MME 550 |
|
|
|
|
76 | 1.6 M Sodium citrate tribasic dihydrate pH 6.5 |
|
|
77 | 0.5 M Ammonium sulfate | 0.1 M HEPES pH 7.5 | 30% v/v MPD |
78 | 0.1 M HEPES pH 7.5 | 10% w/v PEG 6000 | 5% v/v MPD |
79 | 0.1 M HEPES pH 7.5 | 20% v/v Jeffamine M-600 |
|
80 | 0.1 M NaCl | 0.1 M HEPES pH 7.5 | 1.6 M Ammonium sulfate |
81 | 2.0 M Ammonium formate | 0.1 M HEPES pH 7.5 |
|
82 | 0.05 M Cadmium sulfate hydrate | 0.1 M HEPES pH 7.5 | 1.0 M Sodium acetate trihydrate |
83 | 0.1 M HEPES pH 7.5 | 70% v/v MPD |
|
84 | 0.1 M HEPES pH 7.5 | 4.3 M NaCl |
|
85 | 0.1 M HEPES pH 7.5 | 8% v/v Ethylene glycol | 10% w/v PEG 8000 |
86 | 0.1 M HEPES pH 7.5 | 20% w/v PEG 10 | 000 |
87 | 0.2 M MgCl2·6H2O | 0.1 M Tris pH 8.5 | 3.4 M 1, 6-Hexanediol |
88 | 0.1 M Tris pH 8.5 | 25% v/v tert-Butanol |
|
89 | 0.01 M Nickel(II) chloride hexahydrate | 0.1 M Tris pH 8.5 | 1.0 M Lithium sulfate monohydrate |
90 | 1.5 M Ammonium sulfate | 0.1 M Tris pH 8.5 | 12% v/v Glycerol |
91 | 0.2 M Ammonium phosphate monobasic | 0.1 M Tris pH 8.5 | 50% v/v MPD |
92 | 0.1 M Tris pH 8.5 | 20% v/v Ethanol |
|
93 | 0.01 M Nickel(II) chloride hexahydrate | 0.1 M Tris pH 8.5 | 20% w/v PEG MME 2000 |
94 | 0.1 M NaCl | 0.1 M Bicine pH 9.0 | 20% v/v PEG MME 550 |
95 | 0.1 M Bicine pH 9.0 | 2.0 M MgCl2·6H2O |
|
96 | 0.1 M Bicine pH 9.0 | 2% v/v 1, 4-Dioxane | 10% w/v PEG 20000 |
MBP-Fusion Protein Crystallization Screen
Salt/additive | Buffer | Precipitant | |
1 | 0.2 M MgCl2 | MES 6.0 | 20% PEG6000 |
2 | 0.2 M CaCl2 | MES 6.0 | 20% PEG6000 |
3 | 0.2 M Zn acetate | MES 6.0 | 20% PEG6000 |
4 | 0.2 M Li acetate | cacodylate 6.5 | 20% PEG6000 |
5 | 0.2 M Li sulfate | MES 6.0 | 15% PEG6000 |
6 | 0.2 M Na tartrate | MES 6.0 | 20% PEG6000 |
7 | 0.1 M Na citrate | MES 6.0 | 30% PEG6000 |
8 | 0.2 M NH4 acetate | MES 6.0 | 20% PEG6000 |
9 | 0.2 M NH4 sulfate | Na acetate 4.6 | 15% PEG6000 |
10 | 0.2 M NH4 tartrate | MES 6.0 | 20% PEG6000 |
11 | 0.1 M Na/K phosphate | MES 6.0 | 20% PEG6000 |
12 | 0.1 M NaCl, 10 mM MnCl2 | cacodylate 6.5 | 20% PEG6000 |
13 | 0.2 M Na acetate | CHES 9.5 | 20% PEG6000 |
14 | 0.2 M MgCl2 | HEPES 7.4 | 20% PEG6000 |
15 | 0.2 M CaCl2 | HEPES 7.4 | 20% PEG6000 |
16 | 0.2 M Zn acetate | HEPES 6.5 | 20% PEG6000 |
17 | 0.2 M Li acetate | HEPES 7.4 | 25% PEG6000 |
18 | 0.2 M Li sulfate | HEPES 7.4 | 20% PEG6000 |
19 | 0.2 M Na tartrate | HEPES 7.4 | 20% PEG6000 |
20 | 0.2 M Na citrate | HEPES 7.4 | 15% PEG6000 |
21 | 0.2 M NH4 acetate | HEPES 7.4 | 20% PEG6000 |
22 | 0.2 M NH4 sulfate | cacodylate 6.5 | 20% PEG6000 |
23 | 0.2 M NH4 tartrate | HEPES 7.4 | 20% PEG6000 |
24 | 0.1 M Na/K phosphate | HEPES 7.4 | 20% PEG6000 |
25 | 0.1 M Na iodide | Na acetate 4.6 | 20% PEG8000 |
26 | 0.2 M Na citrate | MES 6.0 | 20% PEG8000 |
27 | 0.2 M Na malonate | cacodylate 6.5 | 20% PEG8000 |
28 | 0.2 M NH4 sulfate | MES 6.5 | 20% PEG8000 |
29 | 0.1 M Mg formate | HEPES 7.0 | 20% PEG8000 |
30 | HEPES 7.4 | 20% PEG8000 | |
31 | 0.1 M K nitrate | Tris 8.0 | 20% PEG8000 |
32 | 0.1 M NH4 formate | CHES pH 9.5 | 20% PEG8000 |
33 | 0.1 M Zn acetate | cacodylate 6.5 | 20% PEG8000 |
34 | 0.2 M NH4Cl | Na citrate 5.6 | 15% PEG8000 |
35 | 0.2 M NH4 acetate | Na acetate 5.0 | 15% PEG8000 |
36 | 0.2 M Na/K tartrate | MES 5.0 | 15% PEG8000 |
37 | 0.1 M LiCl | Tris 8.5 | 15% PEG8000 |
38 | 0.1 M MgCl2 | Tris 8.5 | 25% PEG4000 |
39 | 0.2 M NH4 NO3 | Na acetate pH 4.6 | 25% PEG4000 |
40 | 0.1 M NaCl, 10% glycerol | cacodylate 6.5 | 25% PEG4000 |
41 | 0.2 M Na malonate | Tris 8.0 | 25% PEG4000 |
42 | 0.2 M Li sulfate | MES 6.0 | 20% PEG4000 |
43 | 0.2 M Na tartrate | HEPES 7.0 | 20% PEG4000 |
44 | 0.2 M Na formate | HEPES 7.4 | 20% PEG4000 |
45 | 5% isopropynol | Tris 8.5 | 15% PEG4000 |
46 | 0.1 M Zn acetate | cacodylate 7.0 | 15% PEG4000 |
47 | 0.2 Li acetate | Na citrate 5.6 | 15% PEG4000 |
48 | HEPES 7.4 | 20% PEG10000 | |
49 | cacodylate 6.5 | 15% PEG10000 | |
50 | Na citrate 5.6 | 10% PEG10000 | |
51 | 0.2 M NH4 citrate | HEPES 7.0 | 10% PEG10000 |
52 | 0.2 M K acetate | Tris 8.0 | 10% PEG10000 |
53 | 10% glycerol | Na citrate 5.6 | 2.5 M NH4 sulfate |
54 | MES 6.0 | 2.5 M NH4 sulfate | |
55 | HEPES 7.0 | 2.5 M NH4 sulfate | |
56 | Tris 8.0 | 2.5 M NH4 sulfate | |
57 | Na acetate 4.6 | 2.0 M NH4 sulfate | |
58 | MES 5.5 | 2.0 M NH4 sulfate | |
59 | cacodylate 6.5 | 2.0 M NH4 sulfate | |
60 | HEPES 7.4 | 2.0 M NH4 sulfate | |
61 | Tris 8.5 | 2.0 M NH4 sulfate | |
62 | 0.2 M MgCl2 | Na citrate 5.6 | 1.5 M NH4 sulfate |
63 | 10 mM Zn acetate | MES 6.0 | 1.5 M NH4 sulfate |
64 | 10 mM MnCl2 | HEPES 7.0 | 1.5 M NH4 sulfate |
65 | 10 mM CaCl2 | Tris 8.0 | 1.5 M NH4 sulfate |
66 | 1.8 M NaCl, 30% sucrose | ||
67 | 50 mM Cd Cl2 | HEPES 7.4 | 1.0 M Na acetate |
68 | 0.2 M NH4 sulfate | MES 6.5 | 1.0 M Na acetate |
69 | 0.2 M Na acetate, 10 mM MgCl2 | MES 6.0 | 20% PEG5000 MME |
70 | 0.2 M NH4 tartrate | phosphate 7.9 | 20% PEG2000 MME |
71 | 0.2 KCl | Na citrate 5.6 | 20% PEG2000 MME |
72 | 0.2 M NH4 sulfate | Na acetate 4.6 | 20% PEG2000 MME |
73 | 0.2 M NH4 nitrate | cacodylate 6.5 | 20% PEG2000 MME |
74 | 0.1 M NaCl | Na acetate 4.6 | 20% PEG2000 MME |
75 | 0.1 M NaF | Tris 8.5 | 20% PEG2000 MME |
76 | 0.1 M Na/K phosphate | HEPES 7.0 | 20% PEG1000 |
77 | 0.2 M Na malonate | MES 6.0 | 20% PEG1000 |
78 | 0.2 M NH4 sulfate | Na citrate 5.6 | 30% PEG550 MME |
79 | 0.2 M NH4 sulfate | Na acetate 4.6 | 30% PEG550 MME |
80 | 0.1 M MgCl2 | cacodylate 6.5 | 30% PEG550 MME |
81 | 0.1 M KCl | Na acetate 4.6 | 30% PEG550 MME |
82 | Tris 8.5 | 30% PEG550 MME | |
83 | 10 mM CaCl2 | HEPES 7.4 | 35% PEG550 MME |
84 | 10 mM Zn acetate, 0.1 M NaCl | Na citrate 5.6 | 30% PEG400 |
85 | 0.1 M MgCl2 | cacodylate 6.5 | 30% PEG400 |
86 | 0.2 M ammonium sulfate | HEPES 7.0 | 25% PEG400, 25% glycerol |
87 | Na citrate 5.6 | 20% PEG400, 20% sucrose | |
88 | 2% MPD, 50 mM MnCl2 | MES 6.5 | 40% PEG400 |
89 | 50 mM MgCl2 | HEPES 7.4 | 40% PEG400 |
90 | 50 mM CaCl2 | Na acetate 4.6 | 40% PEG400 |
91 | 0.1 M CaCl2 | HEPES 7.0 | 30% MPD |
92 | 0.1 M NaCl | Na citrate 5.6 | 30% MPD |
93 | 0.2 M NH4 sulfate | cacodylate 6.5 | 25% MPD |
94 | 0.1 M K nitrate | MES 5.5 | 20% MPD |
95 | 0.1 M Na/K phosphate | Na acetate 4.6 | 20% MPD |
96 | 10 mM MgCl2, 0.5% Jeffamine | cacodylate 6.0 | 1M Li sulfate |
No. | salts | Buffer | Precipitant |
1 | 0.2 M Sodium fluoride | 20% w/v PEG 3350 | |
2 | 0.2 M Potassium fluoride | 20% w/v PEG 3350 | |
3 | 0.2 M Ammonium fluoride | 20% w/v PEG 3350 | |
4 | 0.2 M Lithium chloride | 20% w/v PEG 3350 | |
5 | 0.2 M Magnesium chloride hexahydrate | 20% w/v PEG 3350 | |
6 | 0.2 M Sodium chloride | 20% w/v PEG 3350 | |
7 | 0.2 M Calcium chloride dihydrate | 20% w/v PEG 3350 | |
8 | 0.2 M Potassium chloride | 20% w/v PEG 3350 | |
9 | 0.2 M Ammonium chloride | 20% w/v PEG 3350 | |
10 | 0.2 M Sodium iodide | 20% w/v PEG 3350 | |
11 | 0.2 M Potassium iodide | 20% w/v PEG 3350 | |
12 | 0.2 M Ammonium iodide | 20% w/v PEG 3350 | |
13 | 0.2 M Sodium thiocyanate | 20% w/v PEG 3350 | |
14 | 0.2 M Potassium thiocyanate | 20% w/v PEG 3350 | |
15 | 0.2 M Lithium nitrate | 20% w/v PEG 3350 | |
16 | 0.2 M Magnesium nitrate hexahydrate | 20% w/v PEG 3350 | |
17 | 0.2 M Sodium nitrate | 20% w/v PEG 3350 | |
18 | 0.2 M Potassium nitrate | 20% w/v PEG 3350 | |
19 | 0.2 M Ammonium nitrate | 20% w/v PEG 3350 | |
20 | 0.2 M Magnesium formate dihydrate | 20% w/v PEG 3350 | |
21 | 0.2 M Sodium formate | 20% w/v PEG 3350 | |
22 | 0.2 M Potassium formate | 20% w/v PEG 3350 | |
23 | 0.2 M Ammonium formate | 20% w/v PEG 3350 | |
24 | 0.2 M Lithium acetate dihydrate | 20% w/v PEG 3350 | |
25 | 0.2 M Magnesium acetate tetrahydrate | 20% w/v PEG 3350 | |
26 | 0.2 M Zinc acetate dihydrate | 20% w/v PEG 3350 | |
27 | 0.2 M Sodium acetate trihydrate | 20% w/v PEG 3350 | |
28 | 0.2 M Calcium acetate hydrate | 20% w/v PEG 3350 | |
29 | 0.2 M Potassium acetate | 20% w/v PEG 3350 | |
30 | 0.2 M Ammonium acetate | 20% w/v PEG 3350 | |
31 | 0.2 M Lithium sulfate monohydrate | 20% w/v PEG 3350 | |
32 | 0.2 M Magnesium sulfate heptahydrate | 20% w/v PEG 3350 | |
33 | 0.2 M Sodium sulfate decahydrate | 20% w/v PEG 3350 | |
34 | 0.2 M Potassium sulfate | 20% w/v PEG 3350 | |
35 | 0.2 M Ammonium sulfate | 20% w/v PEG 3350 | |
36 | 0.2 M Sodium tartrate dibasic dihydrate | 20% w/v PEG 3350 | |
37 | 0.2 M Potassium sodium tartrate | 20% w/v PEG 3350 | |
38 | 0.2 M Ammonium tartrate dibasic | 20% w/v PEG 3350 | |
39 | 0.2 M Sodium dihydrogen phosphate monohydrate | 20% w/v PEG 3350 | |
40 | 0.2 M Sodium phosphate dibasic dihydrate | 20% w/v PEG 3350 | |
41 | 0.2 M Potassium phosphate monobasic | 20% w/v PEG 3350 |
PEG/Ion Screen (HR2-126 + HR2-098)
42 | 0.2 M Potassium phosphate dibasic |
| 20% w/v PEG 3350 |
43 | 0.2 M Ammonium phosphate monobasic |
| 20% w/v PEG 3350 |
44 | 0.2 M Ammonium phosphate dibasic |
| 20% w/v PEG 3350 |
45 | 0.2 M Lithium citrate tribasic tetrahydrate |
| 20% w/v PEG 3350 |
46 | 0.2 M Sodium citrate tribasic dihydrate |
| 20% w/v PEG 3350 |
47 | 0.2 M Potassium citrate tribasic |
| 20% w/v PEG 3350 |
48 | 0.2 M Ammonium citrate dibasic |
| 20% w/v PEG 3350 |
49 | 0.1 M Sodium malonate pH 4.0 | 12% w/v PEG 3350 | |
50 | 0.2 M Sodium malonate pH 4.0 | 20% w/v PEG 3350 | |
51 | 0.1 M Sodium malonate pH 5.0 | 12% w/v PEG 3350 | |
52 | 0.2 M Sodium malonate pH 5.0 | 20% w/v PEG 3350 | |
53 | 0.1 M Sodium malonate pH 6.0 | 12% w/v PEG 3350 | |
54 | 0.2 M Sodium malonate pH 6.0 | 20% w/v PEG 3350 | |
55 | 0.1 M Sodium malonate pH 7.0 | 12% w/v PEG 3350 | |
56 | 0.2 M Sodium malonate pH 7.0 | 20% w/v PEG 3350 | |
57 | 4% v/v Tacsimate pH 4.0 | 12% w/v PEG 3350 | |
58 | 8% v/v Tacsimate pH 4.0 | 20% w/v PEG 3350 | |
59 | 4% v/v Tacsimate pH 5.0 | 12% w/v PEG 3350 | |
60 | 8% v/v Tacsimate pH 5.0 | 20% w/v PEG 3350 | |
61 | 4% v/v Tacsimate pH 6.0 | 12% w/v PEG 3350 | |
62 | 8% v/v Tacsimate pH 6.0 | 20% w/v PEG 3350 | |
63 | 4% v/v Tacsimate pH 7.0 | 12% w/v PEG 3350 | |
64 | 8% v/v Tacsimate pH 7.0 | 20% w/v PEG 3350 | |
65 | 4% v/v Tacsimate pH 8.0 | 12% w/v PEG 3350 | |
66 | 8% v/v Tacsimate pH 8.0 | 20% w/v PEG 3350 | |
67 | 0.1 M Succinic acid pH 7.0 | 12% w/v PEG 3350 | |
68 | 0.2 M Succinic acid pH 7.0 | 20% w/v PEG 3350 | |
69 | 0.1 M Ammonium citrate tribasic pH 7 | 12% w/v PEG 3350 | |
70 | 0.2 M Ammonium citrate tribasic pH 7 | 20% w/v PEG 3350 | |
71 | 0.1 M DL-Malic acid pH 7.0 | 12% w/v PEG 3350 | |
72 | 0.2 M DL-Malic acid pH 7.0 | 20% w/v PEG 3350 | |
73 | 0.1 M Sodium acetate trihydrate pH 7 | 12% w/v PEG 3350 | |
74 | 0.2 M Sodium acetate trihydrate pH 7 | 20% w/v PEG 3350 | |
75 | 0.1 M Sodium formate pH 7.0 | 12% w/v PEG 3350 | |
76 | 0.2 M Sodium formate pH 7.0 | 20% w/v PEG 3350 | |
77 | 0.1 M Ammonium tartrate dibasic pH 7 | 12% w/v PEG 3350 | |
78 | 0.2 M Ammonium tartrate dibasic pH 7 | 20% w/v PEG 3350 | |
79 | 2% v/v Tacsimate pH 4.0 | 0.1 M Sodium acetate trihydrate pH 4.6 | 16% w/v PEG 3350 |
80 | 2% v/v Tacsimate pH 5.0 | 0.1 M Sodium citrate tribasic dihydrate pH 5.6 | 16% w/v PEG 3350 |
81 | 2% v/v Tacsimate pH 6.0 | 0.1 M BIS-TRIS pH 6.5 | 20% w/v PEG 3350 |
PEG/Ion Screen (HR2-126 + HR2-098)
82 | 2% v/v Tacsimate pH 7.0 | 0.1 M HEPES pH 7.5 | 20% w/v PEG 3350 |
83 | 2% v/v Tacsimate pH 8.0 | 0.1 M Tris pH 8.5 | 16% w/v PEG 3350 |
84 | (0.07 M Citric acid | 0.03 M BIS-TRIS propane)/pH 3.4 | 16% w/v PEG 3350 |
85 | (0.06 M Citric acid | 0.04 M BIS-TRIS propane)/pH 4.1 | 16% w/v PEG 3350 |
86 | (0.05 M Citric acid | 0.05 M BIS-TRIS propane)/pH 5.0 | 16% w/v PEG 3350 |
87 | (0.04 M Citric acid | 0.06 M BIS-TRIS propane)/pH 6.4 | 20% w/v PEG 3350 |
88 | (0.03 M Citric acid | 0.07 M BIS-TRIS propane)/pH 7.6 | 20% w/v PEG 3350 |
89 | (0.02 M Citric acid | 0.08 M BIS-TRIS propane)/pH 8.8 | 16% w/v PEG 3350 |
90 | 0.02 M Calcium chloride dihydrate, 0.02 M Cadmium chloride hydrate, 0.02 M Cobalt (II) chloride hexahydrate | 20% w/v PEG 3350 | |
91 | 0.01 M Magnesium chloride hexahydrate, 0.005 M Nickel (II) chloride hexahydrate | 0.1 M HEPES sodium pH 7.0 | 15% w/v PEG 3350 |
92 | 0.02 M Zinc chloride | 20% w/v PEG 3350 | |
93 | 0.15 M Cesium chloride | 15% w/v PEG 3350 | |
94 | 0.2 M Sodium bromide | 20% w/v PEG 3350 | |
95 | 1% w/v Tryptone, 0.001 M Sodium azide | 0.05 M HEPES sodium pH 7.0 | 12% w/v PEG 3350 |
96 | 1% w/v Tryptone, 0.001 M Sodium azide | 0.05 M HEPES sodium pH 7.0 | 20% w/v PEG 3350 |

